In LiNk-NY/curatedTCGAManu: Presents example analyses using curatedTCGAData, MultiAssayExperiment, and TCGAutils
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>",
cache = TRUE,
out.width = "100%"
)
curatedTCGAManu
Multi-omic integration of public oncology databases in Bioconductor
This repository contains scripts and datasets for the
curatedTCGAData + cBioPortalData manuscript.
Overview
We provide several examples to demonstrate the powerful and flexible analysis
environment provided. These analyses, previously only achievable through a
significant investment of time and bioinformatic training, become straight-
forward analysis exercises provided as vignettes.
Key Points
-
Key objective: To provide flexible, integrated multi-omic representations
of public oncology databases in R/Bioconductor with grealy reduced data
management overhead.
-
Knowledge generated: Our Bioconductor software packages provide a novel
approach to lower barriers to analysis and tool development for the TCGA and
cBioPortal databases.
-
Relevance: Our tools provide flexible, programmatic analysis of hundreds of
fully integrated multi’omic oncology datasets within an ecosystem of multi-omic
analysis tools.
Key Packages
- MultiAssayExperiment
- curatedTCGAData
- TCGAutils
- cBioPortalData
- EnrichmentBrowser
- GSEABenchmarkeR
- RTCGAToolbox
Installation
Until the release of Bioconductor 3.11 (scheduled for April 28, 2020), it is
strongly recommended to use the devel version of Bioconductor. That version
can be installed the
traditional way or by
using the Docker container.
Additionally until the release of Bioconductor 3.11, cBioPortalData must be
installed from GitHub as shown in the following code chunk which installs all
necessary packages either directly or as dependencies. Note that this code chunk
is not evaluated, because installation only needs to be performed once.
BiocManager::install(c("cBioPortalData", "LiNk-NY/curatedTCGAManu"))
Vignette Build
Because of the size of the data, it is recommended that the vignettes be built
individually. If using RStudio, the user can simply open the vignette and press
the knit button. Otherwise, the package can be built completely with vignettes
by doing:
R CMD build curatedTCGAManu
in the command line or
BiocManager::install("Link-NY/curatedTCGAManu", build_vignettes = TRUE)
in R.
Loading packages
library(curatedTCGAData)
library(cBioPortalData)
library(TCGAutils)
library(GenomicDataCommons)
library(rtracklayer)
Note. For clarity, we include library commands within the supplemental code
chunks.
Figures
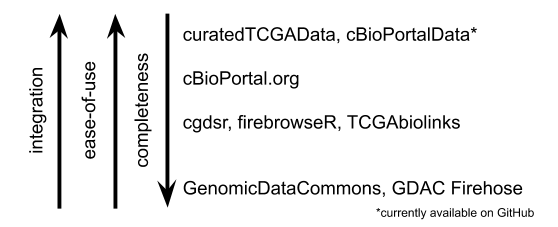
Ease-of-use schematic
Figure 1
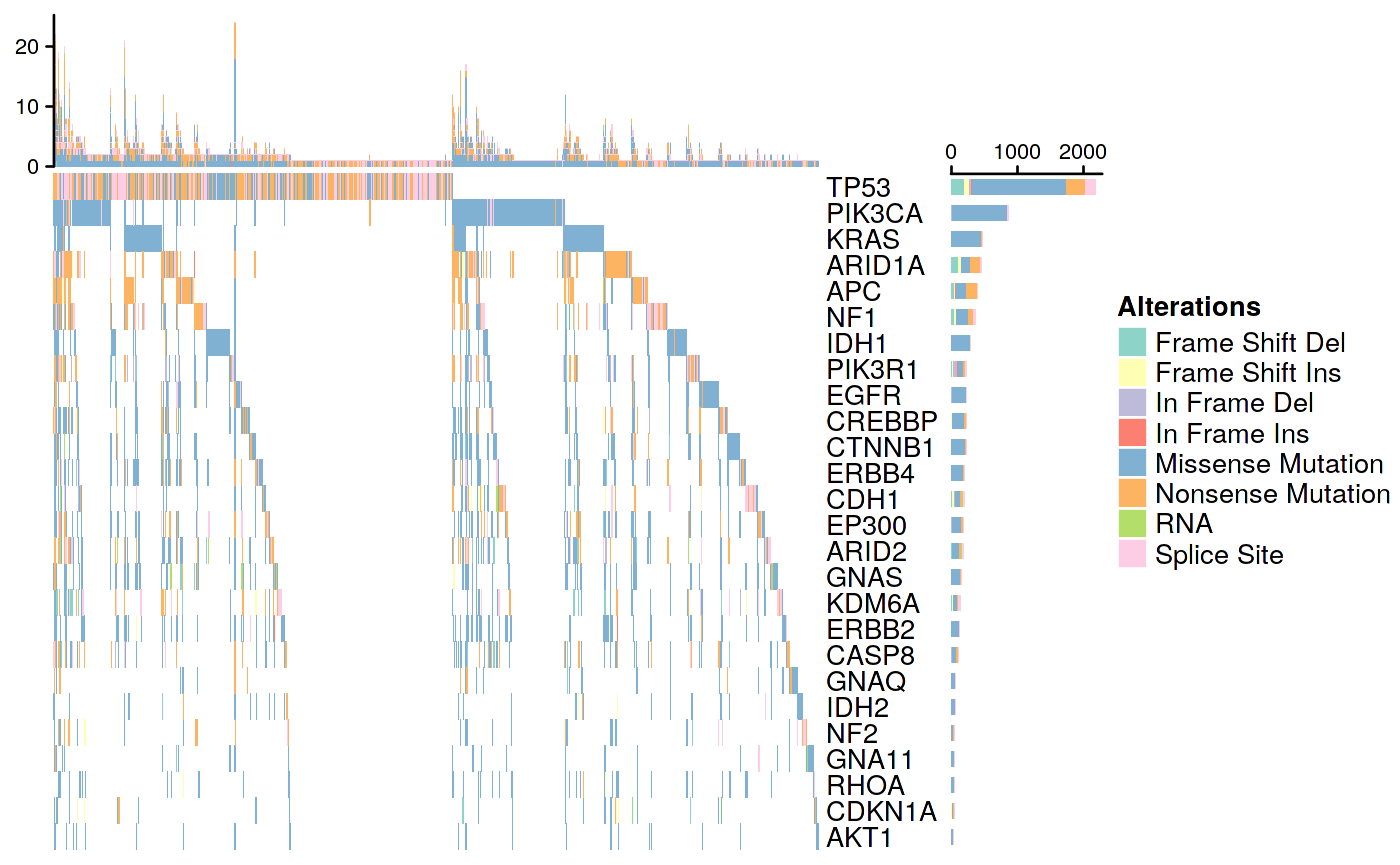
Pan-Cancer OncoPrint Plot
Figure 2
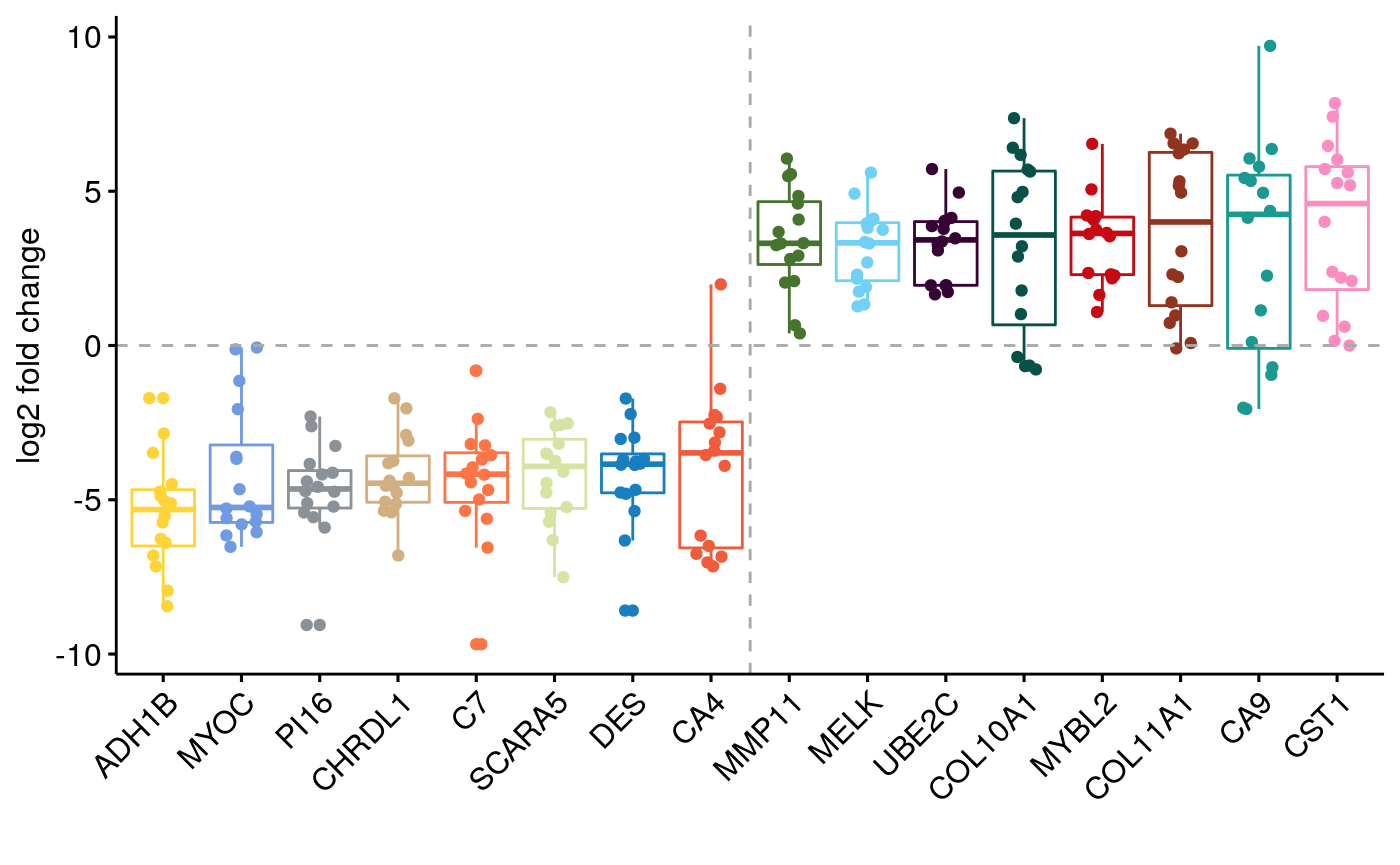
Differential Expression and GSEA PanCan
Figure 3
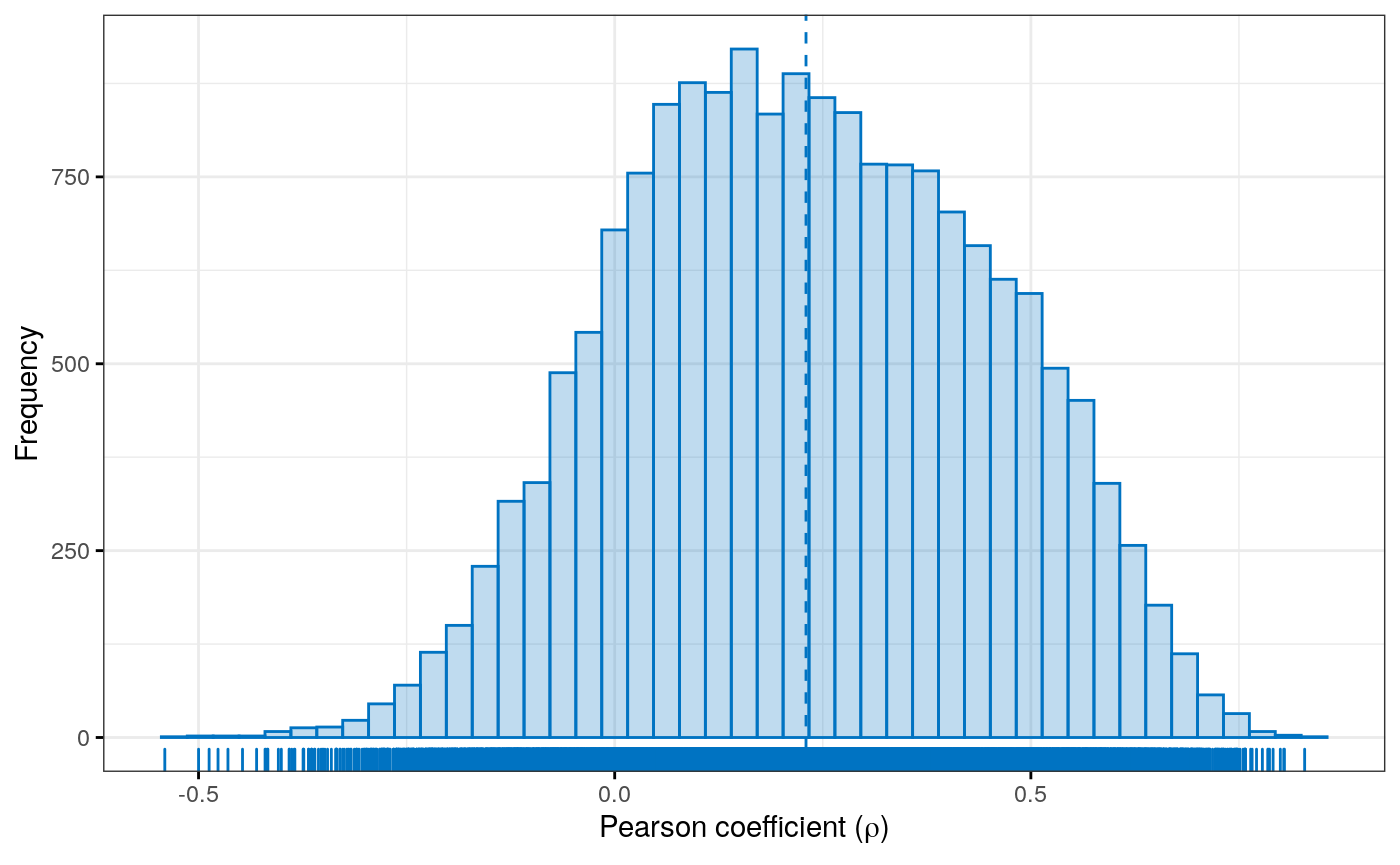
Figure 4
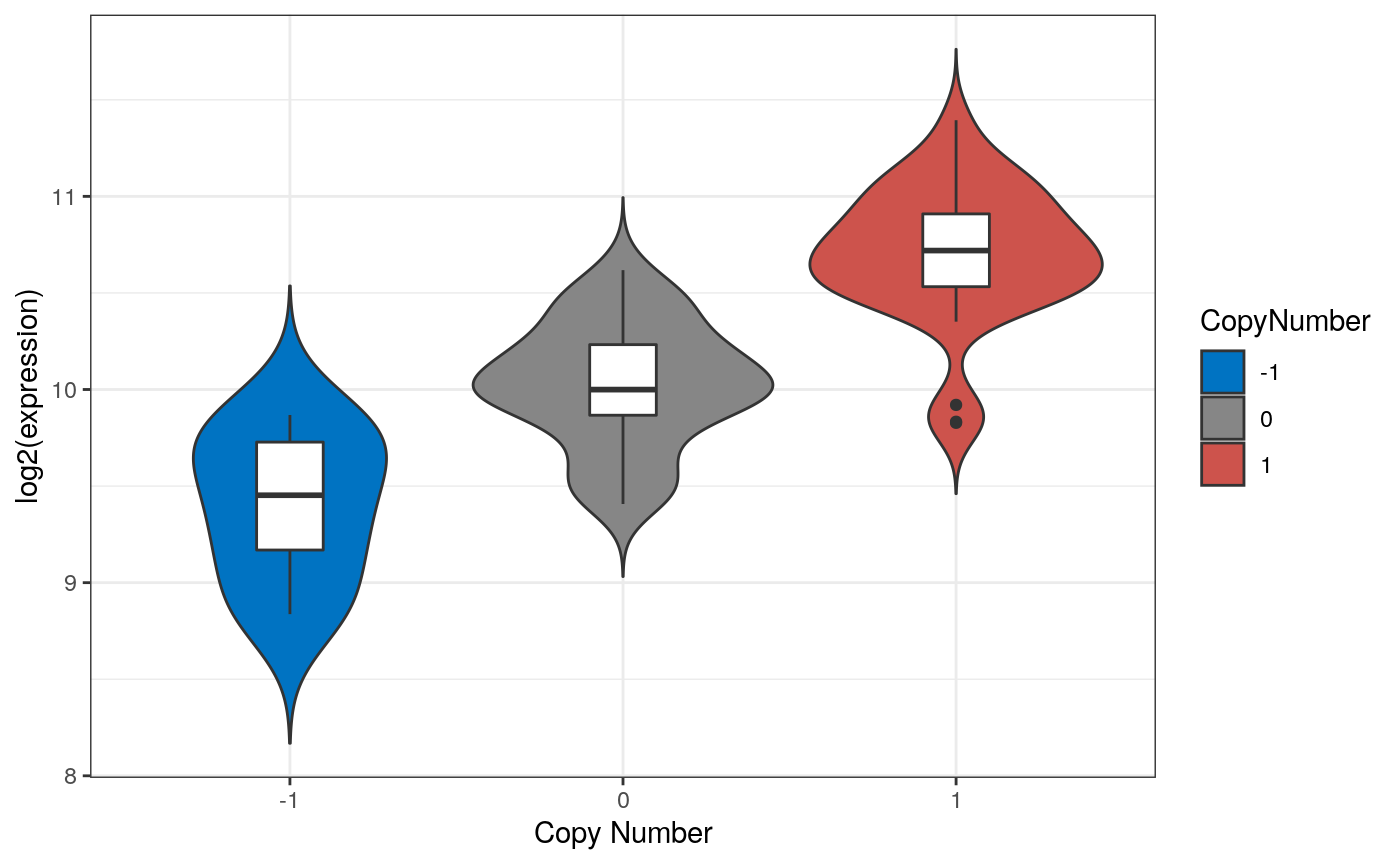
Example Multi-omic Analyses
Figure 5
Figure 6
Supplement Reference
Figure S1
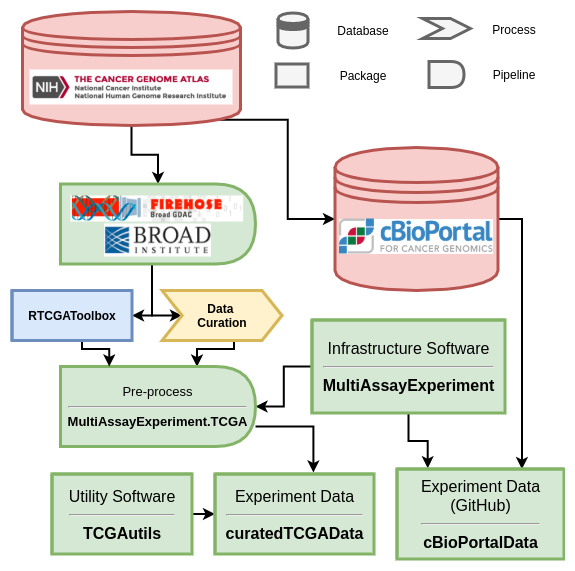
Data provenance and package network
Figure S2A
Example code for installing and downloading TCGA data using curatedTCGAData
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
if (!requireNamespace("curatedTCGAData", quietly = TRUE))
BiocManager::install("curatedTCGAData")
## Glioblastoma Multiforme (GBM)
library(curatedTCGAData)
curatedTCGAData(diseaseCode = "GBM", assays = "RNA*", dry.run = FALSE)
Figure S2B
Example cBioPortalData code for downloading and exporting TCGA data from cBioPortal.org and via the cBioPortal API
## installation
if (!requireNamespace("cBioPortalData", quietly = TRUE))
BiocManager::install("cBioPortalData")
library(cBioPortalData)
## https://cbioportal.org/datasets (Bulk data method)
gbm <- cBioDataPack("gbm_tcga")
## https://cBioPortal.org/api (API method)
cBio <- cBioPortal()
## use exportClass() with the result to save data files
tcga_gbm <- cBioPortalData(cBio, studyId = "gbm_tcga", genePanelId = "IMPACT341")
tcga_gbm
exportClass(tcga_gbm, dir = tempdir(), fmt = "csv")
Figure S2C
Example hg19 to hg38 liftover procedure using Bioconductor tools
liftchain <- "http://hgdownload.cse.ucsc.edu/goldenpath/hg19/liftOver/hg19ToHg38.over.chain.gz"
cloc38 <- file.path(tempdir(), gsub("\\.gz", "", basename(liftchain)))
dfile <- tempfile(fileext = ".gz")
download.file(liftchain, dfile)
R.utils::gunzip(dfile, destname = cloc38, remove = FALSE)
library(rtracklayer)
chain38 <- suppressMessages( import.chain(cloc38) )
## Run bulk data download (from S2B) to create gbm object
if (!exists("gbm")) gbm <- cBioPortalData::cBioDataPack("gbm_tcga")
mutations <- gbm[["mutations_extended"]]
seqlevelsStyle(mutations) <- "UCSC"
ranges38 <- liftOver(rowRanges(mutations), chain38)
Figure S3
Example code for downloading data via GenomicDataCommons and loading with TCGAutils
library(TCGAutils)
library(GenomicDataCommons)
## GenomicDataCommons
query <- files(legacy = TRUE) %>%
filter( ~ cases.project.project_id == "TCGA-COAD" &
data_category == "Gene expression" &
data_type == "Exon quantification" )
fileids <- manifest(query)$id[1:4]
exonfiles <- gdcdata(fileids, use_cached = FALSE)
## TCGAutils
makeGRangesListFromExonFiles(exonfiles, nrows = 4)
Figure S4
Correlated principal components across experimental assays in adrenocortical carcinoma (ACC)
LiNk-NY/curatedTCGAManu documentation built on July 22, 2020, 4:06 p.m.
knitr::opts_chunk$set( collapse = TRUE, comment = "#>", cache = TRUE, out.width = "100%" )
curatedTCGAManu
Multi-omic integration of public oncology databases in Bioconductor
This repository contains scripts and datasets for the curatedTCGAData + cBioPortalData manuscript.
Overview
We provide several examples to demonstrate the powerful and flexible analysis environment provided. These analyses, previously only achievable through a significant investment of time and bioinformatic training, become straight- forward analysis exercises provided as vignettes.
Key Points
-
Key objective: To provide flexible, integrated multi-omic representations of public oncology databases in R/Bioconductor with grealy reduced data management overhead.
-
Knowledge generated: Our Bioconductor software packages provide a novel approach to lower barriers to analysis and tool development for the TCGA and cBioPortal databases.
-
Relevance: Our tools provide flexible, programmatic analysis of hundreds of fully integrated multi’omic oncology datasets within an ecosystem of multi-omic analysis tools.
Key Packages
- MultiAssayExperiment
- curatedTCGAData
- TCGAutils
- cBioPortalData
- EnrichmentBrowser
- GSEABenchmarkeR
- RTCGAToolbox
Installation
Until the release of Bioconductor 3.11 (scheduled for April 28, 2020), it is strongly recommended to use the devel version of Bioconductor. That version can be installed the traditional way or by using the Docker container.
Additionally until the release of Bioconductor 3.11, cBioPortalData must be
installed from GitHub as shown in the following code chunk which installs all
necessary packages either directly or as dependencies. Note that this code chunk
is not evaluated, because installation only needs to be performed once.
BiocManager::install(c("cBioPortalData", "LiNk-NY/curatedTCGAManu"))
Vignette Build
Because of the size of the data, it is recommended that the vignettes be built
individually. If using RStudio, the user can simply open the vignette and press
the knit button. Otherwise, the package can be built completely with vignettes
by doing:
R CMD build curatedTCGAManu
in the command line or
BiocManager::install("Link-NY/curatedTCGAManu", build_vignettes = TRUE)
in R.
Loading packages
library(curatedTCGAData) library(cBioPortalData) library(TCGAutils) library(GenomicDataCommons) library(rtracklayer)
Note. For clarity, we include library commands within the supplemental code
chunks.
Figures
Ease-of-use schematic
Figure 1
Pan-Cancer OncoPrint Plot
Figure 2
Differential Expression and GSEA PanCan
Figure 3
Figure 4
Example Multi-omic Analyses
Figure 5
Figure 6
Supplement Reference
Figure S1
Data provenance and package network
Figure S2A
Example code for installing and downloading TCGA data using curatedTCGAData
if (!requireNamespace("BiocManager", quietly = TRUE)) install.packages("BiocManager") if (!requireNamespace("curatedTCGAData", quietly = TRUE)) BiocManager::install("curatedTCGAData") ## Glioblastoma Multiforme (GBM) library(curatedTCGAData) curatedTCGAData(diseaseCode = "GBM", assays = "RNA*", dry.run = FALSE)
Figure S2B
Example cBioPortalData code for downloading and exporting TCGA data from cBioPortal.org and via the cBioPortal API
## installation if (!requireNamespace("cBioPortalData", quietly = TRUE)) BiocManager::install("cBioPortalData") library(cBioPortalData) ## https://cbioportal.org/datasets (Bulk data method) gbm <- cBioDataPack("gbm_tcga") ## https://cBioPortal.org/api (API method) cBio <- cBioPortal() ## use exportClass() with the result to save data files tcga_gbm <- cBioPortalData(cBio, studyId = "gbm_tcga", genePanelId = "IMPACT341") tcga_gbm exportClass(tcga_gbm, dir = tempdir(), fmt = "csv")
Figure S2C
Example hg19 to hg38 liftover procedure using Bioconductor tools
liftchain <- "http://hgdownload.cse.ucsc.edu/goldenpath/hg19/liftOver/hg19ToHg38.over.chain.gz" cloc38 <- file.path(tempdir(), gsub("\\.gz", "", basename(liftchain))) dfile <- tempfile(fileext = ".gz") download.file(liftchain, dfile) R.utils::gunzip(dfile, destname = cloc38, remove = FALSE) library(rtracklayer) chain38 <- suppressMessages( import.chain(cloc38) ) ## Run bulk data download (from S2B) to create gbm object if (!exists("gbm")) gbm <- cBioPortalData::cBioDataPack("gbm_tcga") mutations <- gbm[["mutations_extended"]] seqlevelsStyle(mutations) <- "UCSC" ranges38 <- liftOver(rowRanges(mutations), chain38)
Figure S3
Example code for downloading data via GenomicDataCommons and loading with TCGAutils
library(TCGAutils) library(GenomicDataCommons) ## GenomicDataCommons query <- files(legacy = TRUE) %>% filter( ~ cases.project.project_id == "TCGA-COAD" & data_category == "Gene expression" & data_type == "Exon quantification" ) fileids <- manifest(query)$id[1:4] exonfiles <- gdcdata(fileids, use_cached = FALSE) ## TCGAutils makeGRangesListFromExonFiles(exonfiles, nrows = 4)
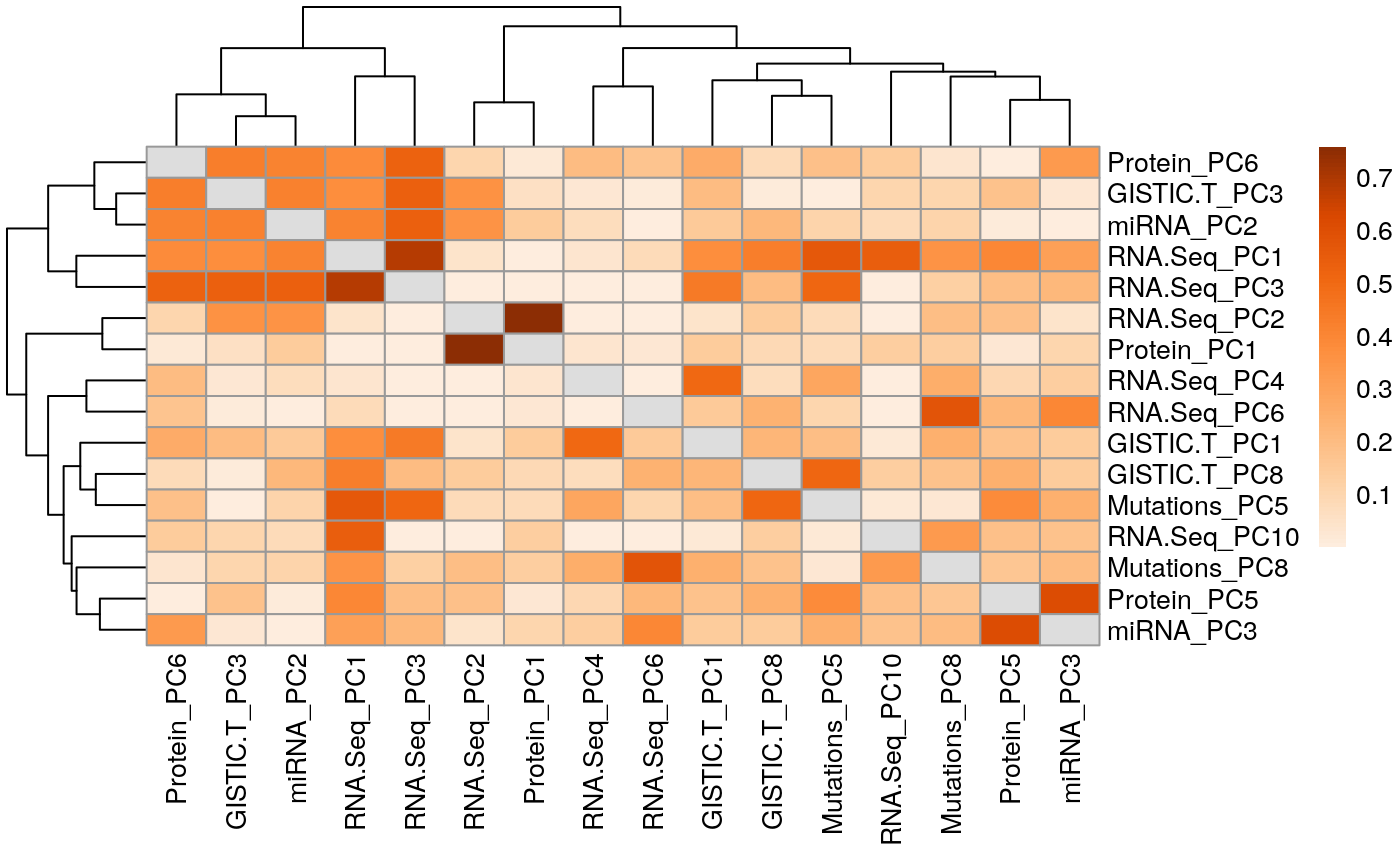
Figure S4
Correlated principal components across experimental assays in adrenocortical carcinoma (ACC)
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.