README.md
In dynamicSDM: Species Distribution and Abundance Modelling at High Spatio-Temporal Resolution
dynamicSDM
Added features dynamicSDM v1.3
-
Added function extract_static_coords for extracting
spatially-buffered co-ordinate data from static datasets.
-
Added arguments to dynamic_proj_covariates() for adding static
rasters to covariates for each data (e.g. static elevation raster)
-
Removed package dependency on raster, sp, geodist and
geosphere.
-
All functions are now terra and sf compatible.
-
The package has since been published in the Open Access journal
“Methods in Ecology and Evolution”
Dobson, R., Challinor, A.J., Cheke, R.A., Jennings, S., Willis, S.G.
and Dallimer, M., 2023. dynamicSDM: An R package for species
geographical distribution and abundance modelling at high spatiotemporal
resolution. Methods in Ecology and Evolution, 14, 1190-1199.
Summary
Across ecological research fields, species distribution and abundance
modelling (SDM) is a major tool for understanding the drivers and
patterns of species occurrence. To advance our ability to model species
inhabiting dynamic ecosystems worldwide, dynamicSDM facilitates the
incorporation of explanatory variables that are dynamic in both space
and time. Our functions are:
- user-friendly - requiring only simple inputs and outputs;
- highly flexible - offering diverse and open arguments for
targeting study specifics;
- computer friendly - utilising Google Earth Engine and Google Drive
to minimise the computing power and storage demands associated with
high spatiotemporal resolution modelling.
Package structure
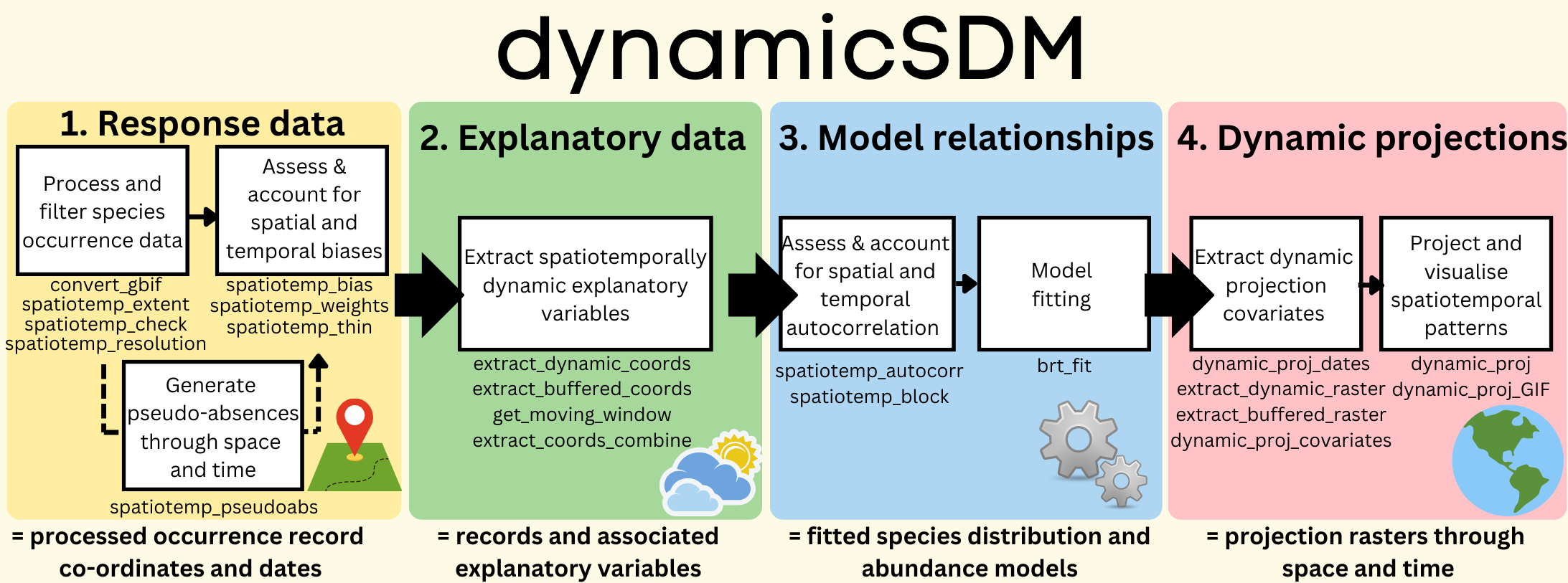
dynamicSDM functions are split into four key modelling stages: response
data, explanatory variables, modelling relationships and dynamic
projections. See the package manual
here
for more details on each function.
1) Response data functions
Functions for preparing species distribution or abundance model input
data for modelling with spatiotemporally dynamic explanatory variables.
convert_gbif() Transform Global Biodiversity Information Facility
occurrence records to dynamicSDM compatible.spatiotemp_check() Check species occurrence record formatting,
completeness and validity.spatiotemp_extent() Filter species occurrence records by a given
spatial and temporal extent.spatiotemp_resolution() Filter species occurrence records by given
spatial and temporal resolution.spatiotemp_bias() Test for spatial and temporal bias in species
occurrence records.spatiotemp_thin() Thin species occurrence records by spatial and
temporal proximity.spatiotemp_pseudoabs() Generate pseudo-absence record coordinates
and dates.spatiotemp_weights() Calculate sampling effort across spatial and
temporal buffer from occurrence records.
2) Explanatory variable functions
Functions for extracting spatiotemporally dynamic explanatory variable
data for species occurrence record co-ordinates and dates using Google
Earth Engine.
extract_dynamic_coords() Extract temporally dynamic explanatory
variable data for occurrence records.get_moving_window() Generate a “moving window” matrix of optimal
size for spatial buffering of explanatory variable data.extract_buffered_coords() Extract spatially buffered and temporally
dynamic explanatory variable data for occurrence records.extract_coords_combine() Combine extracted explanatory variable data
for occurrence records into single data frame for model fitting.extract_static_coords Extract spatially buffered data from static
rasters for occurrence record co-ordinates (no temporal dimension).
3) Modelling relationship functions
Functions for generating species distribution or abundance models that
account for spatial and temporal autocorrelation in dynamic explanatory
variables.
spatiotemp_autocorr() Test for spatial and temporal autocorrelation
in species distribution model explanatory data.spatiotemp_block() Split occurrence records into spatial and
temporal blocks for model fitting.brt_fit() Fit boosted regression tree models to species distribution
or abundance data.
4) Dynamic projection functions
Functions for generating explanatory variable projection data frames at
given spatiotemporal extent and resolution, and projecting species
dynamic distribution and abundance patterns onto these.
dynamic_proj_dates() Generate vector of dates for dynamic
projectionsextract_dynamic_raster() Extract temporally dynamic rasters of
explanatory variable data.extract_buffered_raster() Extract spatially buffered and temporally
dynamic rasters of explanatory variable data.dynamic_proj_covariates() Combine explanatory variable rasters into
a covariate data frame for each projection date.dynamic_proj() Project species distribution and abundance models
onto dynamic environmental covariates.dynamic_proj_GIF() Create GIF of dynamic species distribution and
abundance projections
Installation
# Install using Github
install_github("r-a-dobson/dynamicSDM")
Common installation errors
dynamicSDM depends on a range of spatial and graphic R packages, which
may result in some persistent errors on installation or running of
certain functions.
If you encounter an error or bug when installing and using dynamicSDM,
please post a comment
here for guidance and
support from us.
Below we have outlined common errors and typical solutions to try,
depending on your operating system
1) Error with rgl
# Loading rgl's DLL failed. This build of rgl depends on XQuartz, which failed to load.
options(rgl.useNULL = TRUE)
library(rgl)
2) Dependency package terra
On Homebrew (macOS) run:
brew install pkg-config
brew install gdal
On Linux run:
sudo apt-get install libgdal-dev libproj-dev libgeos-dev libudunits2-dev netcdf-bin
Then in R run:
install.packages("Rcpp")
install.packages('terra', repos='https://rspatial.r-universe.dev')
3) Dependency package magick
On Homebrew (macOS) run:
brew install imagemagick@6
On Linux run:
sudo apt-get install -y libmagick++-dev
Then in R run:
install.packages("magick")
Try the dynamicSDM package in your browser
Any scripts or data that you put into this service are public.
dynamicSDM documentation built on June 28, 2024, 5:08 p.m.
dynamicSDM
Added features dynamicSDM v1.3
-
Added function
extract_static_coordsfor extracting spatially-buffered co-ordinate data from static datasets. -
Added arguments to
dynamic_proj_covariates()for adding static rasters to covariates for each data (e.g. static elevation raster) -
Removed package dependency on
raster,sp,geodistandgeosphere. -
All functions are now
terraandsfcompatible. -
The package has since been published in the Open Access journal “Methods in Ecology and Evolution”
Dobson, R., Challinor, A.J., Cheke, R.A., Jennings, S., Willis, S.G. and Dallimer, M., 2023. dynamicSDM: An R package for species geographical distribution and abundance modelling at high spatiotemporal resolution. Methods in Ecology and Evolution, 14, 1190-1199.
Summary
Across ecological research fields, species distribution and abundance modelling (SDM) is a major tool for understanding the drivers and patterns of species occurrence. To advance our ability to model species inhabiting dynamic ecosystems worldwide, dynamicSDM facilitates the incorporation of explanatory variables that are dynamic in both space and time. Our functions are:
- user-friendly - requiring only simple inputs and outputs;
- highly flexible - offering diverse and open arguments for targeting study specifics;
- computer friendly - utilising Google Earth Engine and Google Drive to minimise the computing power and storage demands associated with high spatiotemporal resolution modelling.
Package structure
dynamicSDM functions are split into four key modelling stages: response data, explanatory variables, modelling relationships and dynamic projections. See the package manual here for more details on each function.
1) Response data functions
Functions for preparing species distribution or abundance model input data for modelling with spatiotemporally dynamic explanatory variables.
convert_gbif()Transform Global Biodiversity Information Facility occurrence records todynamicSDMcompatible.spatiotemp_check()Check species occurrence record formatting, completeness and validity.spatiotemp_extent()Filter species occurrence records by a given spatial and temporal extent.spatiotemp_resolution()Filter species occurrence records by given spatial and temporal resolution.spatiotemp_bias()Test for spatial and temporal bias in species occurrence records.spatiotemp_thin()Thin species occurrence records by spatial and temporal proximity.spatiotemp_pseudoabs()Generate pseudo-absence record coordinates and dates.spatiotemp_weights()Calculate sampling effort across spatial and temporal buffer from occurrence records.
2) Explanatory variable functions
Functions for extracting spatiotemporally dynamic explanatory variable data for species occurrence record co-ordinates and dates using Google Earth Engine.
extract_dynamic_coords()Extract temporally dynamic explanatory variable data for occurrence records.get_moving_window()Generate a “moving window” matrix of optimal size for spatial buffering of explanatory variable data.extract_buffered_coords()Extract spatially buffered and temporally dynamic explanatory variable data for occurrence records.extract_coords_combine()Combine extracted explanatory variable data for occurrence records into single data frame for model fitting.extract_static_coordsExtract spatially buffered data from static rasters for occurrence record co-ordinates (no temporal dimension).
3) Modelling relationship functions
Functions for generating species distribution or abundance models that account for spatial and temporal autocorrelation in dynamic explanatory variables.
spatiotemp_autocorr()Test for spatial and temporal autocorrelation in species distribution model explanatory data.spatiotemp_block()Split occurrence records into spatial and temporal blocks for model fitting.brt_fit()Fit boosted regression tree models to species distribution or abundance data.
4) Dynamic projection functions
Functions for generating explanatory variable projection data frames at given spatiotemporal extent and resolution, and projecting species dynamic distribution and abundance patterns onto these.
dynamic_proj_dates()Generate vector of dates for dynamic projectionsextract_dynamic_raster()Extract temporally dynamic rasters of explanatory variable data.extract_buffered_raster()Extract spatially buffered and temporally dynamic rasters of explanatory variable data.dynamic_proj_covariates()Combine explanatory variable rasters into a covariate data frame for each projection date.dynamic_proj()Project species distribution and abundance models onto dynamic environmental covariates.dynamic_proj_GIF()Create GIF of dynamic species distribution and abundance projections
Installation
# Install using Github
install_github("r-a-dobson/dynamicSDM")
Common installation errors
dynamicSDM depends on a range of spatial and graphic R packages, which may result in some persistent errors on installation or running of certain functions.
If you encounter an error or bug when installing and using dynamicSDM, please post a comment here for guidance and support from us.
Below we have outlined common errors and typical solutions to try, depending on your operating system
1) Error with rgl
# Loading rgl's DLL failed. This build of rgl depends on XQuartz, which failed to load.
options(rgl.useNULL = TRUE)
library(rgl)
2) Dependency package terra
On Homebrew (macOS) run:
brew install pkg-config
brew install gdal
On Linux run:
sudo apt-get install libgdal-dev libproj-dev libgeos-dev libudunits2-dev netcdf-bin
Then in R run:
install.packages("Rcpp")
install.packages('terra', repos='https://rspatial.r-universe.dev')
3) Dependency package magick
On Homebrew (macOS) run:
brew install imagemagick@6
On Linux run:
sudo apt-get install -y libmagick++-dev
Then in R run:
install.packages("magick")
Try the dynamicSDM package in your browser
Any scripts or data that you put into this service are public.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.