README.md
In Benli11/rPCA: Randomized Singular Value Decomposition

Fast Randomized Singular Value Decomposition using R
Randomized singular value decomposition (rsvd) is a fast probabilistic algorithm that can
be used to compute the near optimal low-rank singular value decomposition of massive data sets with high accuracy.
The key idea is to compute a compressed representation
of the data to capture the essential information. This compressed representation can then be used to obtain
the low-rank singular value decomposition decomposition. The rsvd package provides one of the fastest routines for low-rank matrix approximations in R, as far as we know.
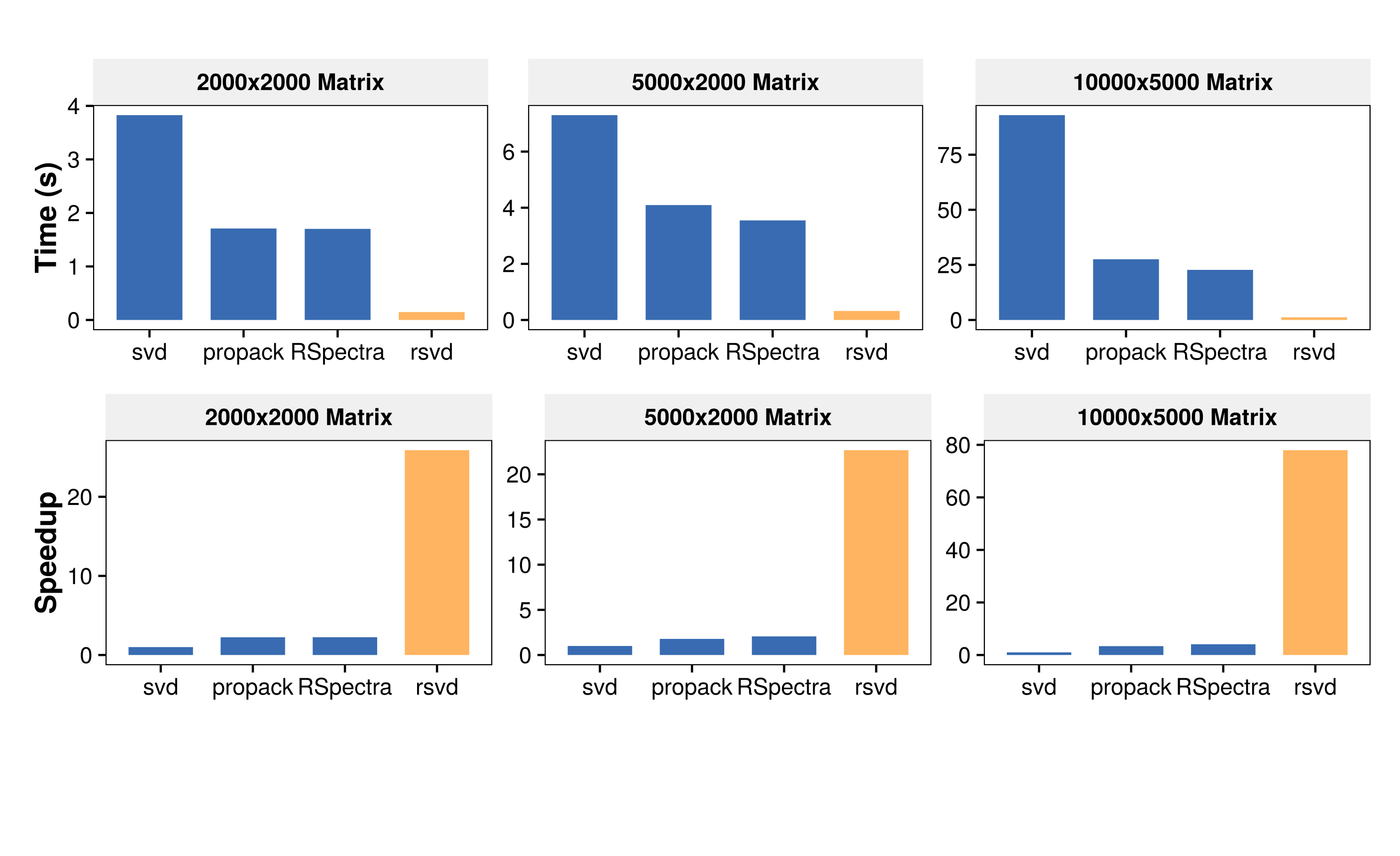
The computational advantage becomes pronounced with an increasing matrix dimension (here target-rank k=50):
The singular value decomposition plays a central role in data analysis and scientific computing.
The SVD is also widely used for computing
(randomized) principal component analysis (PCA), a linear dimensionality reduction technique.
Randomized PCA (rpca) uses the approximated singular value decomposition
to compute the most significant principal components. This package also includes a
function to compute (randomized) robust principal component analysis (RPCA).
In addition several plot functions are provided. See for further details: Randomized Matrix Decompositions using R.
SVD example: Image compression
library(rsvd)
data(tiger)
# Image compression using randomized SVD
s <- rsvd(tiger, k=150)
tiger.re = s$u %*% diag(s$d) %*% t(s$v) # reconstruct image
# Display orginal and reconstrucuted image
par(mfrow=c(1,2))
image(tiger, col = gray((0:255)/255))
image(tiger.re, col = gray((0:255)/255))
Here are the results:
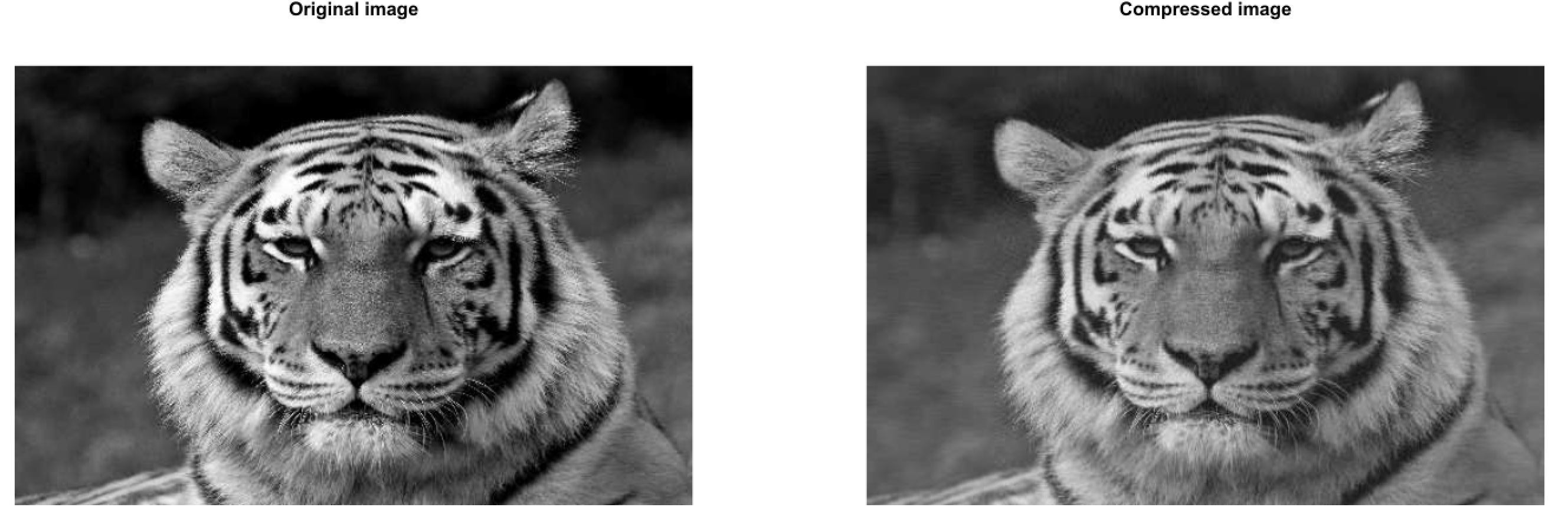

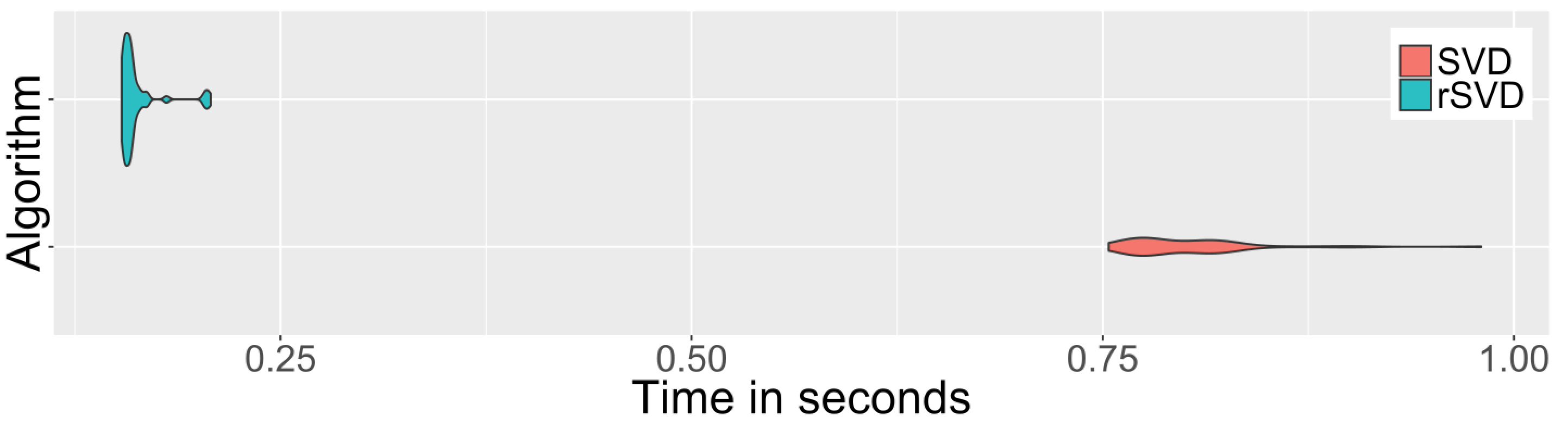
and the speedup gained over the base SVD function:
library(microbenchmark)
timing_svd <- microbenchmark(
'SVD' = svd(tiger, nu=150, nv=150),
'rSVD' = rsvd(tiger, k=150),
times=50)
print(timing_svd, unit='s')
Installation
Install the rsvd package via CRAN
install.packages("rsvd")
You can also install the development version from GitHub using devtools:
devtools::install_github("erichson/rsvd")
The source packge can be obtained here: CRAN: rsvd.
New in Version 1.0.2
- Several small issues are fixed.
- Thanks to Aaron Lun, who has fixed a bug in the rsvd function that occured when nu=0 or nv=0.
New in Version 1.0.0
- Support for non-default matrix types to deal with large-scale matrices that are held on file, added by Aaron Lun.
- Fixed a bug which occured runninig rpca with k=1 and retx=TRUE, discovered by Will.
References
- Erichson NB, Voronin S, Brunton SL, Kutz JN (2019). Randomized Matrix Decompositions Using R. Journal of Statistical Software, 89(11), 1–48. doi: 10.18637/jss.v089.i11.
- Sergey Voronin, Per-Gunnar Martinsson. RSVDPACK: Subroutines for computing partial singular value decompositions via randomized sampling on single core, multi core, and GPU architectures. (2015)
- Nathan Halko, et al. Finding structure with randomness: Probabilistic algorithms for constructing approximate matrix decompositions. (2011)
Cite as
@Article{,
title = {Randomized Matrix Decompositions Using {R}},
author = {N. Benjamin Erichson and Sergey Voronin and Steven L.
Brunton and J. Nathan Kutz},
journal = {Journal of Statistical Software},
year = {2019},
volume = {89},
number = {11},
pages = {1--48},
doi = {10.18637/jss.v089.i11},
}
Benli11/rPCA documentation built on April 20, 2021, 6:50 a.m.

Fast Randomized Singular Value Decomposition using R
Randomized singular value decomposition (rsvd) is a fast probabilistic algorithm that can be used to compute the near optimal low-rank singular value decomposition of massive data sets with high accuracy. The key idea is to compute a compressed representation of the data to capture the essential information. This compressed representation can then be used to obtain the low-rank singular value decomposition decomposition. The rsvd package provides one of the fastest routines for low-rank matrix approximations in R, as far as we know. The computational advantage becomes pronounced with an increasing matrix dimension (here target-rank k=50):

The singular value decomposition plays a central role in data analysis and scientific computing. The SVD is also widely used for computing (randomized) principal component analysis (PCA), a linear dimensionality reduction technique. Randomized PCA (rpca) uses the approximated singular value decomposition to compute the most significant principal components. This package also includes a function to compute (randomized) robust principal component analysis (RPCA). In addition several plot functions are provided. See for further details: Randomized Matrix Decompositions using R.
SVD example: Image compression
library(rsvd)
data(tiger)
# Image compression using randomized SVD
s <- rsvd(tiger, k=150)
tiger.re = s$u %*% diag(s$d) %*% t(s$v) # reconstruct image
# Display orginal and reconstrucuted image
par(mfrow=c(1,2))
image(tiger, col = gray((0:255)/255))
image(tiger.re, col = gray((0:255)/255))
Here are the results:

and the speedup gained over the base SVD function:
library(microbenchmark)
timing_svd <- microbenchmark(
'SVD' = svd(tiger, nu=150, nv=150),
'rSVD' = rsvd(tiger, k=150),
times=50)
print(timing_svd, unit='s')

Installation
Install the rsvd package via CRAN
install.packages("rsvd")
You can also install the development version from GitHub using devtools:
devtools::install_github("erichson/rsvd")
The source packge can be obtained here: CRAN: rsvd.
New in Version 1.0.2
- Several small issues are fixed.
- Thanks to Aaron Lun, who has fixed a bug in the rsvd function that occured when nu=0 or nv=0.
New in Version 1.0.0
- Support for non-default matrix types to deal with large-scale matrices that are held on file, added by Aaron Lun.
- Fixed a bug which occured runninig rpca with k=1 and retx=TRUE, discovered by Will.
References
- Erichson NB, Voronin S, Brunton SL, Kutz JN (2019). Randomized Matrix Decompositions Using R. Journal of Statistical Software, 89(11), 1–48. doi: 10.18637/jss.v089.i11.
- Sergey Voronin, Per-Gunnar Martinsson. RSVDPACK: Subroutines for computing partial singular value decompositions via randomized sampling on single core, multi core, and GPU architectures. (2015)
- Nathan Halko, et al. Finding structure with randomness: Probabilistic algorithms for constructing approximate matrix decompositions. (2011)
Cite as
@Article{,
title = {Randomized Matrix Decompositions Using {R}},
author = {N. Benjamin Erichson and Sergey Voronin and Steven L.
Brunton and J. Nathan Kutz},
journal = {Journal of Statistical Software},
year = {2019},
volume = {89},
number = {11},
pages = {1--48},
doi = {10.18637/jss.v089.i11},
}
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
