In GordonChengTW/GeneticAlgorithm:
針對航空公司主要的收入來源—求取最佳獲利能力
dataSource <- data.frame(
seat = 1000,
cost = 100000,
price = c(100, 300, 600, 800, 3000),
rate = c(0.9, 0.85, 0.7, 0.6, 0.3),
weight= c(0.5,1,1,1,0.5)
);
config <- list(
iters = 60,
population.size = 30,
chromosome.size = length(dataSource$price),
mutate.rate = 0.5
);
- Initial Population
- 平均分散法
- 預期乘坐率
population.initial <- function(
) {
# TODO : implement initialization function/interface
chromSize <- config$chromosome.size;
seat <- as.integer(dataSource$seat / chromSize);
result <- rep(0, chromSize);
for (i in 1:chromSize) {
rate <- dataSource$rate[i] + 1.0;
seatNum <- as.integer(seat * rate);
result[i] <- sample(0:seatNum, size=1);
}
return (result);
}
chromosome.fitness <- function(chromosome
) {
# TODO : implement fintness function/interface
chromSize <- config$chromosome.size;
chrom <- as.vector(chromosome);
seatNum <- sum(chrom);
overNum <- as.logical(seatNum > dataSource$seat);
values <- rep(0, chromSize);
#
C <- as.numeric(dataSource$cost);
for (i in 1:chromSize) {
Q <- as.numeric(chromosome[i]);
P <- as.numeric(dataSource$price[i]);
S <- as.double((dataSource$rate[i]) ^ (i*2));
V <- as.integer(as.integer(Q * P - C) * S * dataSource$weight[i]);
#V <- as.integer(Q * P - C);
values[i] <- ifelse(overNum, -abs(V), V);
}
sum(values);
}
-
- chromosomes
-
- 選取兩組最好的組合
- 當前世代
- 祖先世代 (跨代)
population.selection <- function(dataset,
iters,
current=NA,
callback
) {
# TODO : implement selection function/interface
c1 <- NA;
c2 <- NA;
bestObj <- getBestObj(current, function(chrom) {
c1 <<- chrom;
});
getBestObj(dataset, function(chrom) {
c2 <<- chrom;
});
#
if (is.function(callback)){
callback(bestObj);
}
rbind(c1, c2);
}
- Crossover function
- 跨世代隨機選擇半數基因
- 父世代:2組
- 子世代:1組
chromosome.crossover <- function(dataset
) {
# TODO : implement crossover function/interface
parents <- dataset;
rows <- nrow(parents);
cols <- ncol(parents);
num <- as.integer(cols / 2);
child <- rep(0, cols);
#seat <- as.integer(dataSource$seat);
#seat <- as.integer(dataSource$seat / chromSize);
for (i in 1:num) {
pIdx <- sample(1:rows, size=1);
parent <- parents[pIdx, ];
cIdx <- sample(1:cols, size=1);
child[cIdx] <- parent[cIdx];
#weight <- dataSource$weight[i];
#Q1 <- seat * weight;
#Q2 <- parent[i];
#Q3 <- as.integer(mean(Q1, Q2));
#child[i] <- Q3;
}
child;
}
- Mutation function
- 隨機突變:
- 週期突變:2代
- 突變範圍:半數基因
- 突變參數:乘坐率 (not yet)
chromosome.mutation <- function(chromosome,
iters
) {
# TODO : implement mutation function/interface
mutate.rate <- config$mutate.rate;
chrom <- as.vector(chromosome);
if (runif(1,0,1) < mutate.rate || iters %% 2==0) {
cols <- length(chrom);
count <- as.integer(cols / 2);
seatNum <- as.integer(dataSource$seat / cols);
for (i in 1:count) {
p <- sample(1:cols, size=1);
n <- sample(0:1, size=1);
x <- ifelse(n==0, 1, -1);
m <- sample(0:seatNum, size=1);
chrom[p] <- abs(chrom[p] + x * m) # %% seat;
}
}
chrom
}
-
TODO: GA Flowchart
-
演算結束回呼方法
onFinalize <-function(dataset,
lastPopulation=NA,
callback=NA
) {
# TODO : implement finalize function/interface
if (is.function(callback)){
bestObj <- getBestObj(lastPopulation);
callback(bestObj);
}
}
- 獲取最佳物件的實作方法
getBestObj <- function(dataset, callback=NA) {
idx <- which.max(dataset[, 1]);
bestObj <- dataset[idx, ];
if (is.function(callback)){
chrom <- gcga.getChromosome(bestObj);
callback(chrom);
}
return(bestObj);
}
- Evaluate Chromosomes
library("gcga");
GAmodel <- gcga(iters = config$iters,
population.size = config$population.size,
chromosome.size = config$chromosome.size,
onInit = population.initial,
onFitness = chromosome.fitness,
onSelection = population.selection,
onCrossover = chromosome.crossover,
onMutation = chromosome.mutation,
onFinalize = onFinalize);
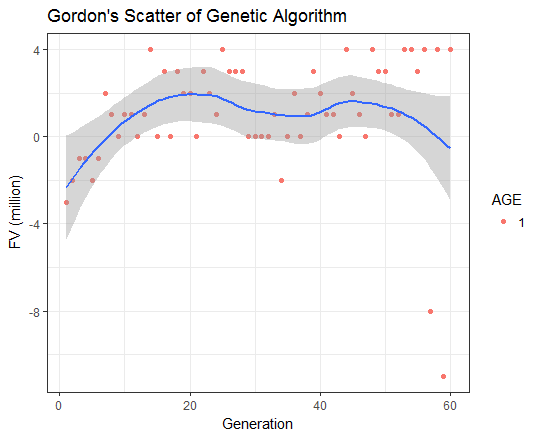
GordonChengTW/GeneticAlgorithm documentation built on May 6, 2019, 6:30 p.m.
針對航空公司主要的收入來源—求取最佳獲利能力
dataSource <- data.frame( seat = 1000, cost = 100000, price = c(100, 300, 600, 800, 3000), rate = c(0.9, 0.85, 0.7, 0.6, 0.3), weight= c(0.5,1,1,1,0.5) ); config <- list( iters = 60, population.size = 30, chromosome.size = length(dataSource$price), mutate.rate = 0.5 );
- Initial Population
- 平均分散法
- 預期乘坐率
population.initial <- function( ) { # TODO : implement initialization function/interface chromSize <- config$chromosome.size; seat <- as.integer(dataSource$seat / chromSize); result <- rep(0, chromSize); for (i in 1:chromSize) { rate <- dataSource$rate[i] + 1.0; seatNum <- as.integer(seat * rate); result[i] <- sample(0:seatNum, size=1); } return (result); }
chromosome.fitness <- function(chromosome ) { # TODO : implement fintness function/interface chromSize <- config$chromosome.size; chrom <- as.vector(chromosome); seatNum <- sum(chrom); overNum <- as.logical(seatNum > dataSource$seat); values <- rep(0, chromSize); # C <- as.numeric(dataSource$cost); for (i in 1:chromSize) { Q <- as.numeric(chromosome[i]); P <- as.numeric(dataSource$price[i]); S <- as.double((dataSource$rate[i]) ^ (i*2)); V <- as.integer(as.integer(Q * P - C) * S * dataSource$weight[i]); #V <- as.integer(Q * P - C); values[i] <- ifelse(overNum, -abs(V), V); } sum(values); }
-
- chromosomes
-
- 選取兩組最好的組合
- 當前世代
- 祖先世代 (跨代)
- 選取兩組最好的組合
population.selection <- function(dataset, iters, current=NA, callback ) { # TODO : implement selection function/interface c1 <- NA; c2 <- NA; bestObj <- getBestObj(current, function(chrom) { c1 <<- chrom; }); getBestObj(dataset, function(chrom) { c2 <<- chrom; }); # if (is.function(callback)){ callback(bestObj); } rbind(c1, c2); }
- Crossover function
- 跨世代隨機選擇半數基因
- 父世代:2組
- 子世代:1組
- 跨世代隨機選擇半數基因
chromosome.crossover <- function(dataset ) { # TODO : implement crossover function/interface parents <- dataset; rows <- nrow(parents); cols <- ncol(parents); num <- as.integer(cols / 2); child <- rep(0, cols); #seat <- as.integer(dataSource$seat); #seat <- as.integer(dataSource$seat / chromSize); for (i in 1:num) { pIdx <- sample(1:rows, size=1); parent <- parents[pIdx, ]; cIdx <- sample(1:cols, size=1); child[cIdx] <- parent[cIdx]; #weight <- dataSource$weight[i]; #Q1 <- seat * weight; #Q2 <- parent[i]; #Q3 <- as.integer(mean(Q1, Q2)); #child[i] <- Q3; } child; }
- Mutation function
- 隨機突變:
- 週期突變:2代
- 突變範圍:半數基因
- 突變參數:乘坐率 (not yet)
chromosome.mutation <- function(chromosome, iters ) { # TODO : implement mutation function/interface mutate.rate <- config$mutate.rate; chrom <- as.vector(chromosome); if (runif(1,0,1) < mutate.rate || iters %% 2==0) { cols <- length(chrom); count <- as.integer(cols / 2); seatNum <- as.integer(dataSource$seat / cols); for (i in 1:count) { p <- sample(1:cols, size=1); n <- sample(0:1, size=1); x <- ifelse(n==0, 1, -1); m <- sample(0:seatNum, size=1); chrom[p] <- abs(chrom[p] + x * m) # %% seat; } } chrom }
-
TODO: GA Flowchart
-
演算結束回呼方法
onFinalize <-function(dataset, lastPopulation=NA, callback=NA ) { # TODO : implement finalize function/interface if (is.function(callback)){ bestObj <- getBestObj(lastPopulation); callback(bestObj); } }
- 獲取最佳物件的實作方法
getBestObj <- function(dataset, callback=NA) { idx <- which.max(dataset[, 1]); bestObj <- dataset[idx, ]; if (is.function(callback)){ chrom <- gcga.getChromosome(bestObj); callback(chrom); } return(bestObj); }
- Evaluate Chromosomes
library("gcga"); GAmodel <- gcga(iters = config$iters, population.size = config$population.size, chromosome.size = config$chromosome.size, onInit = population.initial, onFitness = chromosome.fitness, onSelection = population.selection, onCrossover = chromosome.crossover, onMutation = chromosome.mutation, onFinalize = onFinalize);

Embedding an R snippet on your website
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
