In andrefaa/ENM_TheMetaLand: Create and process Ecological Niche, including several pre- and post-processing methods
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>",
warning=F,
echo = TRUE)
We present here the anwser to some questions that are more common to arise while running ENMTML. If you couldn't find your issue among those questions, please report it and we will make our best to help you run the package smoothly.
At first the number fo arguments and the folder's structure may seem a little scary, but with little time you will get the hang of it and will start running ENM models!
Project Folder
We highly recommend you to create a folder for your project!
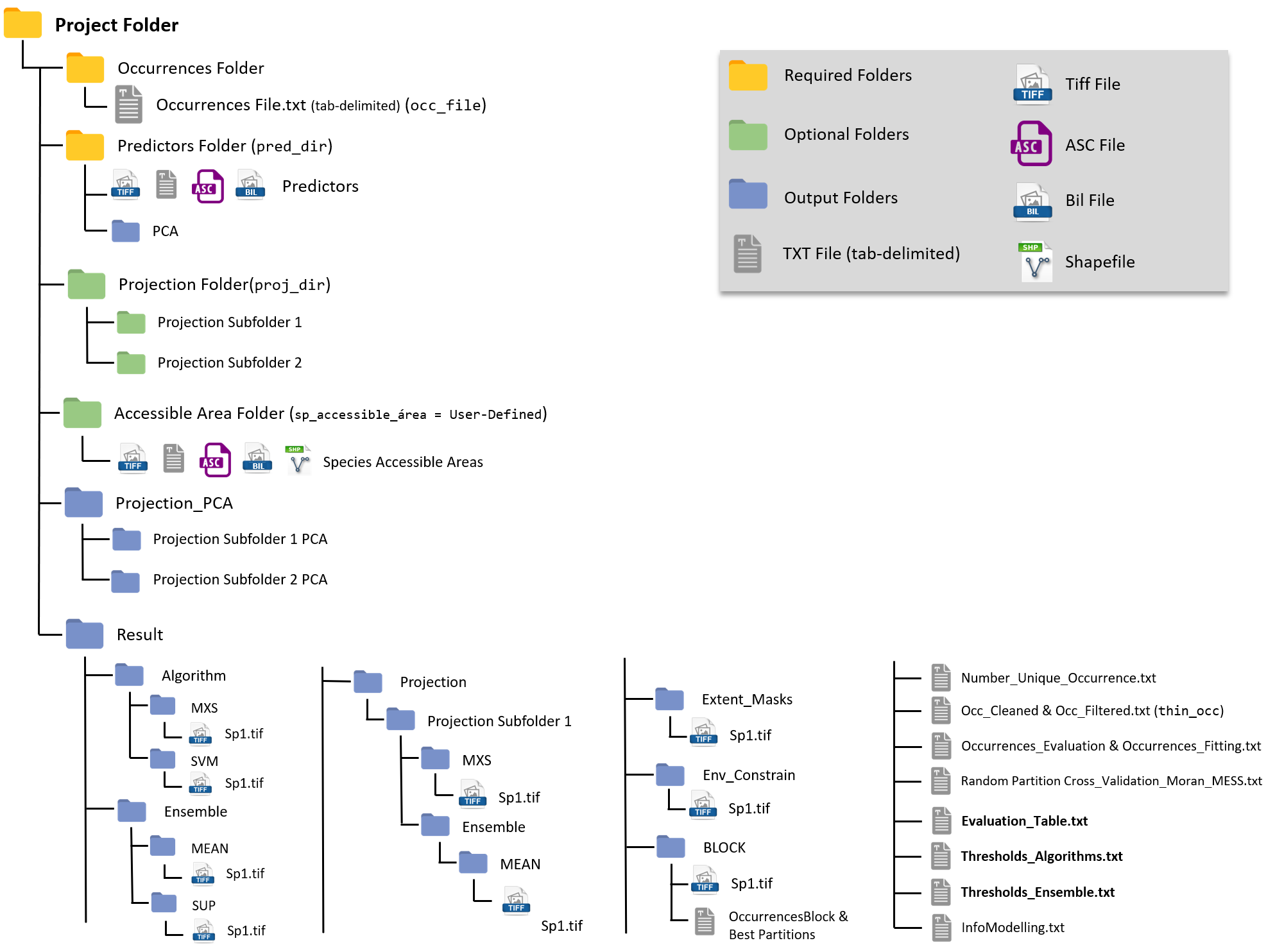
A general overview of the Project Folder and its containing sub-folders will look similar to this:
Yellow are the Mandatory Folders, you need them to run the package.
Green are the Optional Folders, only required according to your project design.
Blue are Output Folders, those will be created during the modeling routine.
Function Arguments
ENMTML consists of several arguments, which will be checked at the start of the modeling routine.
Take your time to check any Error returned during this argument check and use the help to correctly fill the arguments.
Occurrence Data
Occurrence File Type
ENMTML imports occurrence data from a TXT file with tabulation as separator (tab-separated)
Data Format
The TXT can have several columns, but three columns are mandatory:
- Species name
- Longitude
- Latitude
They can be named to your preference as those will be inputted in the arguments sp, x, y of the ENMTML function.
Here's an example of an occurrence table for four sample species.
require(ENMTML)
data(occ)
knitr::kable(occ[c(1:3,43:45,90:92,127:129),],row.names=F)
Predictors
Predictor File Type
ENMTML accepts predictors from several formats: TIF, BIL, ASC and TXT(tab-separated)
Folder Arrangement
Predictors must be within a single folder, which will be informed through the pred_dir argument.
Projections
Projections are indicated using the proj_dir argument.
As it is common to usually make projections to multiple climate scenarios or extents, the arrangement of Projection folders consists in a major folder with different sub-folders (one for each climatic scenario or extent).
It is essential that the name of the variables in those folder match the names of variables within the pred_dir folder!
Results
Results will be in a folder specified in the result_dir argument, or if you chose not to supply any folder ENMTML will create a folder named Results at the same level of your pred_dir folder (see this at the figure at the Project Folder section.
Model Evaluation and information about the modelling routine can be found in several txt files within the Results folder.
Important sub-folders with result maps are:
- Algorithm: with tif present projections for the selected algorithms and the binary folders for the selected thresholds
- Ensemble: with tif projections for the selected ensembles.
- Projection: with tif projections for the different climatic scenarios/extents
Other sub-folders will be created according to the modeling routine and contain masks related to pseudo-absence allocation and geographical partition of datasets.
andrefaa/ENM_TheMetaLand documentation built on Nov. 15, 2023, 10:19 a.m.
knitr::opts_chunk$set( collapse = TRUE, comment = "#>", warning=F, echo = TRUE)
We present here the anwser to some questions that are more common to arise while running ENMTML. If you couldn't find your issue among those questions, please report it and we will make our best to help you run the package smoothly.
At first the number fo arguments and the folder's structure may seem a little scary, but with little time you will get the hang of it and will start running ENM models!
Project Folder
We highly recommend you to create a folder for your project!
A general overview of the Project Folder and its containing sub-folders will look similar to this:
Yellow are the Mandatory Folders, you need them to run the package.
Green are the Optional Folders, only required according to your project design.
Blue are Output Folders, those will be created during the modeling routine.

Function Arguments
ENMTML consists of several arguments, which will be checked at the start of the modeling routine.
Take your time to check any Error returned during this argument check and use the help to correctly fill the arguments.
Occurrence Data
Occurrence File Type
ENMTML imports occurrence data from a TXT file with tabulation as separator (tab-separated)
Data Format
The TXT can have several columns, but three columns are mandatory:
- Species name
- Longitude
- Latitude
They can be named to your preference as those will be inputted in the arguments sp, x, y of the ENMTML function.
Here's an example of an occurrence table for four sample species.
require(ENMTML) data(occ) knitr::kable(occ[c(1:3,43:45,90:92,127:129),],row.names=F)
Predictors
Predictor File Type
ENMTML accepts predictors from several formats: TIF, BIL, ASC and TXT(tab-separated)
Folder Arrangement
Predictors must be within a single folder, which will be informed through the pred_dir argument.
Projections
Projections are indicated using the proj_dir argument.
As it is common to usually make projections to multiple climate scenarios or extents, the arrangement of Projection folders consists in a major folder with different sub-folders (one for each climatic scenario or extent).
It is essential that the name of the variables in those folder match the names of variables within the pred_dir folder!
Results
Results will be in a folder specified in the result_dir argument, or if you chose not to supply any folder ENMTML will create a folder named Results at the same level of your pred_dir folder (see this at the figure at the Project Folder section.
Model Evaluation and information about the modelling routine can be found in several txt files within the Results folder.
Important sub-folders with result maps are:
- Algorithm: with tif present projections for the selected algorithms and the binary folders for the selected thresholds
- Ensemble: with tif projections for the selected ensembles.
- Projection: with tif projections for the different climatic scenarios/extents
Other sub-folders will be created according to the modeling routine and contain masks related to pseudo-absence allocation and geographical partition of datasets.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
