README.md
In celehs/sureLDA: A Novel Multi-Disease Automated Phenotyping Method for the Electronic Health Record
sureLDA: A Novel Multi-Disease Automated Phenotyping Method for the EHR
Overview
Surrogate-guided ensemble Latent Dirichlet Allocation (sureLDA) is a
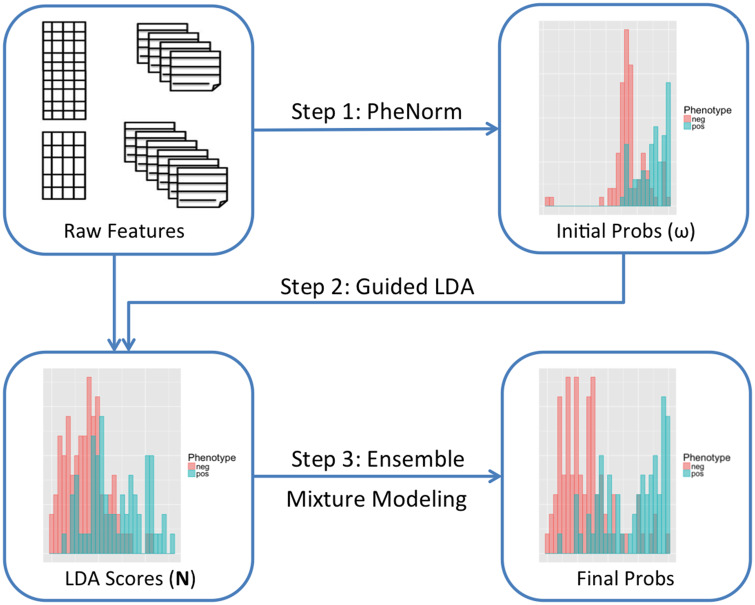
label-free multidimensional phenotyping method. It first uses the
PheNorm or MAP algorithm to initialize probabilities based on two
surrogate features for each target disease, and then leverages these
probabilities to guide the LDA topic model to generate
phenotype-specific topics. Finally, it combines phenotype-feature counts
with surrogates via clustering ensemble to yield final phenotype
probabilities.
Installation
Install stable version from CRAN:
install.packages("sureLDA")
Install development version from GitHub:
# install.packages("remotes")
devtools::install_github("remotes/sureLDA")
Citation
Ahuja Y, Zhou D, He Z, Sun J, Castro VM, Gainer V, Murphy SN, Hong C, Cai T. sureLDA: A multidisease automated phenotyping method for the electronic health record. J Am Med Inform Assoc. 2020 Aug 1;27(8):1235-1243. doi: 10.1093/jamia/ocaa079. PMID: 32548637; PMCID: PMC7481024.
celehs/sureLDA documentation built on June 13, 2025, 6:20 a.m.
sureLDA: A Novel Multi-Disease Automated Phenotyping Method for the EHR
Overview
Surrogate-guided ensemble Latent Dirichlet Allocation (sureLDA) is a label-free multidimensional phenotyping method. It first uses the PheNorm or MAP algorithm to initialize probabilities based on two surrogate features for each target disease, and then leverages these probabilities to guide the LDA topic model to generate phenotype-specific topics. Finally, it combines phenotype-feature counts with surrogates via clustering ensemble to yield final phenotype probabilities.

Installation
Install stable version from CRAN:
install.packages("sureLDA")
Install development version from GitHub:
# install.packages("remotes")
devtools::install_github("remotes/sureLDA")
Citation
Ahuja Y, Zhou D, He Z, Sun J, Castro VM, Gainer V, Murphy SN, Hong C, Cai T. sureLDA: A multidisease automated phenotyping method for the electronic health record. J Am Med Inform Assoc. 2020 Aug 1;27(8):1235-1243. doi: 10.1093/jamia/ocaa079. PMID: 32548637; PMCID: PMC7481024.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
