README.md
In cipheLab/CIPHoLD:
CIPHoLD : Adaptation of the SCAFFOLD cytometry tool.
Install dependencies
To use this tool you need a windows computer with :
- R version 3.5.1 or later
- R studio
- R tools 3.5 (administrator installation and check "Path environment" box during installation)
- Google Chrome browser (to download zip output correctly)
Now run your Rstudio and you can copy paste this commande line to install all apcajges requierd by CIPHoLD tools:
install.packages(c("devtools","shiny","shinydashboard","shinyjs","gtools","shinyHeatmaply"))
install.packages("lme4")
install.packages("BiocManager")
BiocManager::install("flowCore", version = "3.8")
BiocManager::install("ggcyto", version = "3.8")
library("devtools")
install_github("cipheLab/CIPHoLD")
Run Tool
library("CIPHoLD")
CIPHoLD.run()
Presentation
Lexical
- Over-clustering : Its a volontaryto ask more cluster than population expected with unsupervised algorithme. With that you can play like a downsampling data which each cluster is a heterogenous cell groups (its not a population, we don't have this information).This approch it's used in SPADE, FlowSOM and other clustering algorithme
- Enrichment :
- Annotation :
CIPHoLD Tools is a rewrite of SCAFFOLD tools (https://github.com/nolanlab/scaffold).
Difference with scaffold
- File manager
- Upload CSV file (clustering from other tools)
- Add other clustering method
- Write annotation in FCS files
- All interface update with dashboard
- Different output format and information
We decide to change the input and output system.
Upload Data
You can upload FCS in first tab and create group. Group creation is just a concatenation of FCS with new parameters used to save the files id (sample)

upload your multiple fcs file , you can upload 5G of data, if you can't make a issu or send mail and we change this option.
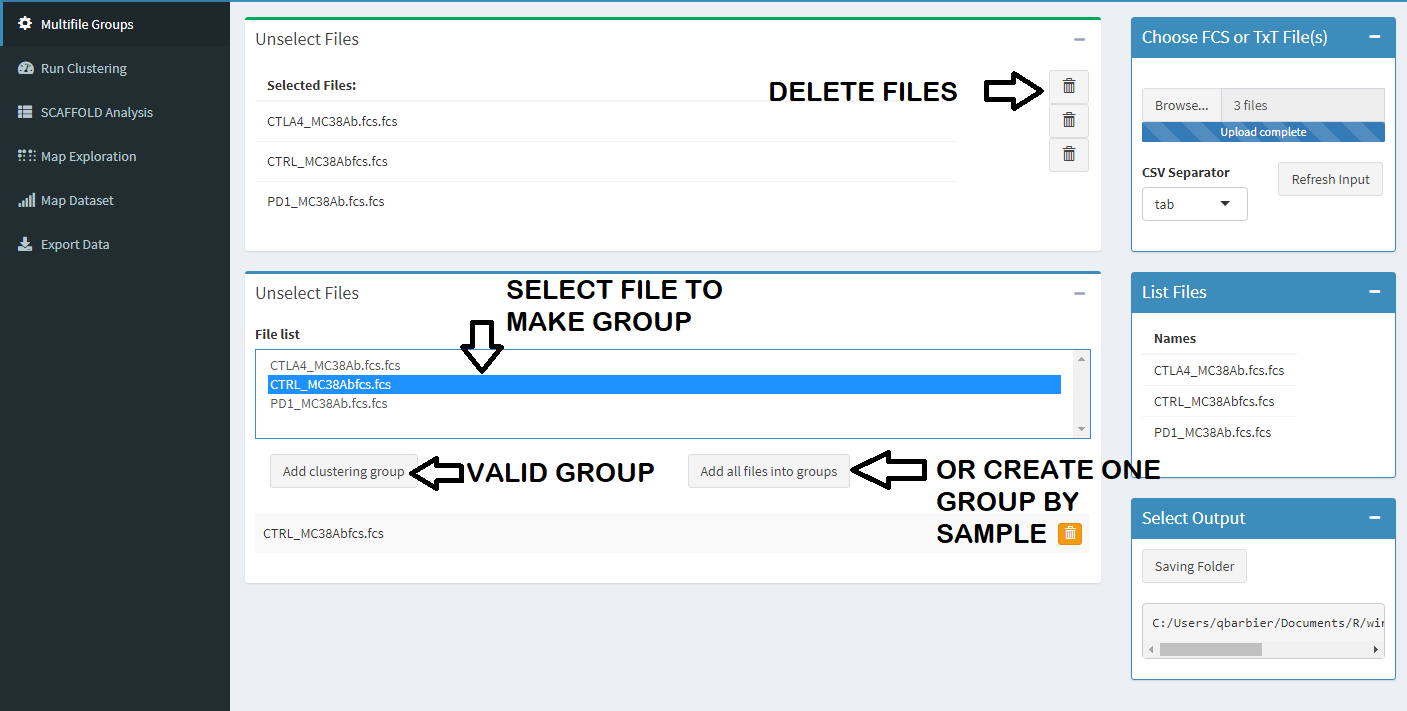
You can create one groupe by file or concantenate multiple file (sample) in one. Its a good idea to donc change the cluster position
in forced-directe-graph and just view a size cluster changing

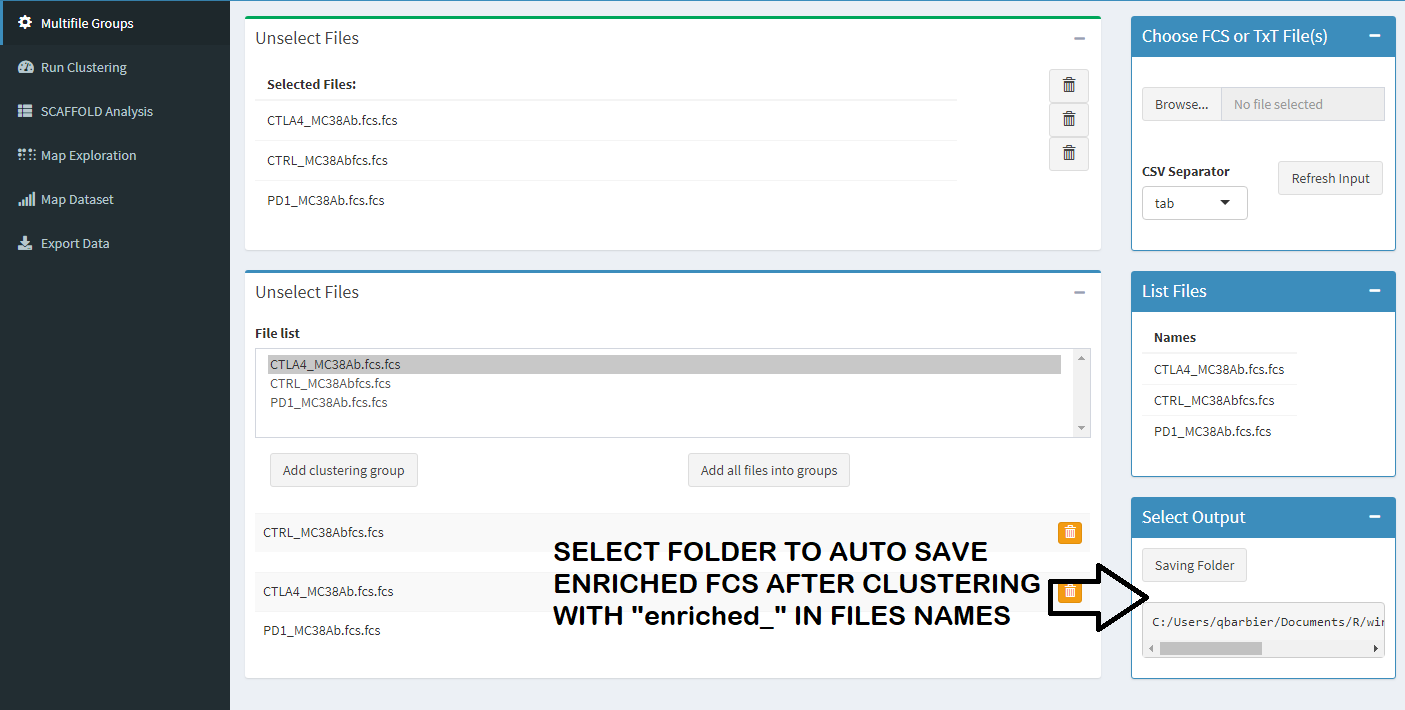
You can upload csv table, with this files you don't need to clustering. Its result of clustering parts
Clustering
The clustering is an over-clustering. You need ask more than 100 clusters to make works the annotation. With CIPHoLD you can use gated file from one technology (flow, mass, rnaseq, etc..) for create map and annotate fcs file from other technology, because you can use different transformation on clustered file and gated files.
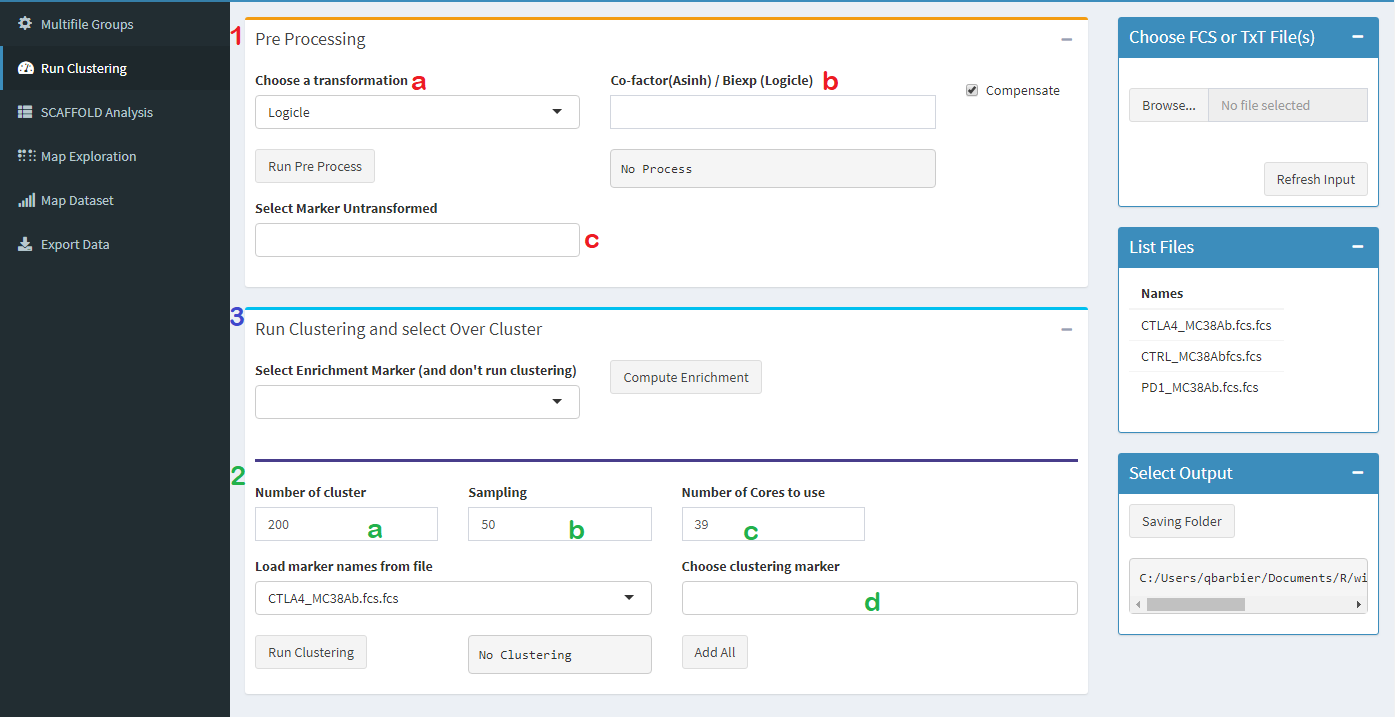
1 : a : Select transform function you want use, arcsinh (masse data with 5 in argument or 150 for flow data) or logicle transform with biexp arg (value below zero save in fcs as $PnW keyword for each parameters
1 : b : Select argument in transform function and compensate or not. If its mass data and they are no "SPILL"keyword automaticly compensate a not engage.
1 : c : If you don't want transform value, like clustering enrichment, annotaion, t-sne coordinate, or other reduction dimensions or information, its possible to "safe" your parameters in this part
2 : a : If you upad enriched fcss wih over-cspas in expression matrce, you need to select this params in this field. Don't forget, if it's necessary, to transform other value and don't transforrm your enrichment parameters
3 : a :
Gated Files and Map creation
Map new dataset
Explore Mapping
Download Annotation
cipheLab/CIPHoLD documentation built on June 12, 2019, 2:43 a.m.
CIPHoLD : Adaptation of the SCAFFOLD cytometry tool.
Install dependencies
To use this tool you need a windows computer with :- R version 3.5.1 or later
- R studio
- R tools 3.5 (administrator installation and check "Path environment" box during installation)
- Google Chrome browser (to download zip output correctly)
Now run your Rstudio and you can copy paste this commande line to install all apcajges requierd by CIPHoLD tools:
install.packages(c("devtools","shiny","shinydashboard","shinyjs","gtools","shinyHeatmaply"))
install.packages("lme4")
install.packages("BiocManager")
BiocManager::install("flowCore", version = "3.8")
BiocManager::install("ggcyto", version = "3.8")
library("devtools")
install_github("cipheLab/CIPHoLD")
Run Tool
library("CIPHoLD")
CIPHoLD.run()
Presentation
Lexical
- Over-clustering : Its a volontaryto ask more cluster than population expected with unsupervised algorithme. With that you can play like a downsampling data which each cluster is a heterogenous cell groups (its not a population, we don't have this information).This approch it's used in SPADE, FlowSOM and other clustering algorithme
- Enrichment :
- Annotation :
CIPHoLD Tools is a rewrite of SCAFFOLD tools (https://github.com/nolanlab/scaffold).
- File manager
- Upload CSV file (clustering from other tools)
- Add other clustering method
- Write annotation in FCS files
- All interface update with dashboard
- Different output format and information
Difference with scaffold
Upload Data
You can upload FCS in first tab and create group. Group creation is just a concatenation of FCS with new parameters used to save the files id (sample)

upload your multiple fcs file , you can upload 5G of data, if you can't make a issu or send mail and we change this option.

You can create one groupe by file or concantenate multiple file (sample) in one. Its a good idea to donc change the cluster position in forced-directe-graph and just view a size cluster changing


You can upload csv table, with this files you don't need to clustering. Its result of clustering parts
Clustering
The clustering is an over-clustering. You need ask more than 100 clusters to make works the annotation. With CIPHoLD you can use gated file from one technology (flow, mass, rnaseq, etc..) for create map and annotate fcs file from other technology, because you can use different transformation on clustered file and gated files.

1 : a : Select transform function you want use, arcsinh (masse data with 5 in argument or 150 for flow data) or logicle transform with biexp arg (value below zero save in fcs as $PnW keyword for each parameters
1 : b : Select argument in transform function and compensate or not. If its mass data and they are no "SPILL"keyword automaticly compensate a not engage.
1 : c : If you don't want transform value, like clustering enrichment, annotaion, t-sne coordinate, or other reduction dimensions or information, its possible to "safe" your parameters in this part
2 : a : If you upad enriched fcss wih over-cspas in expression matrce, you need to select this params in this field. Don't forget, if it's necessary, to transform other value and don't transforrm your enrichment parameters
3 : a :
Gated Files and Map creation
Map new dataset
Explore Mapping
Download Annotation
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
