README.md
In jamez-eh/casc: Casc
casc
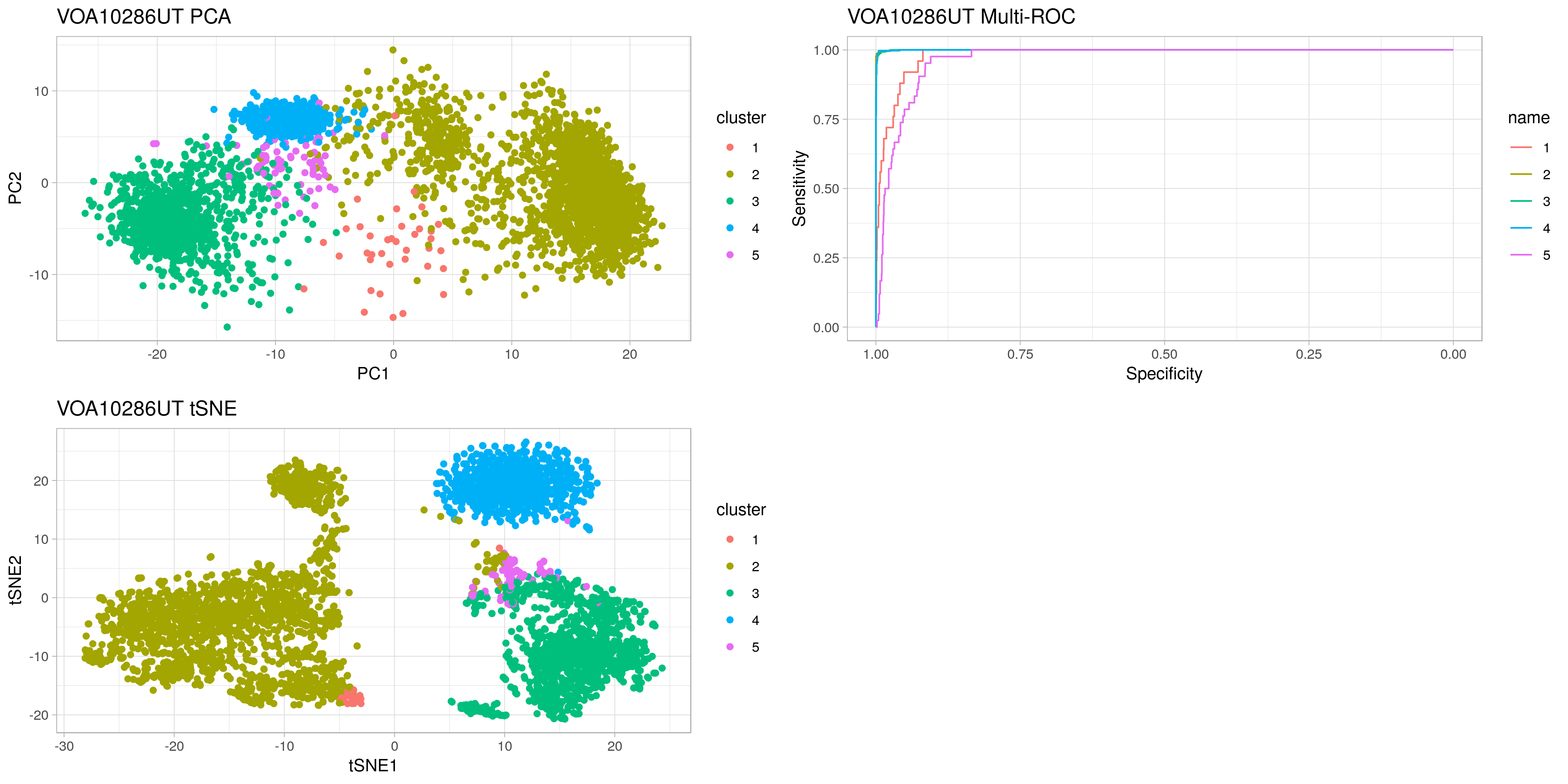
casc trains a logisitic regression model on provided single-cell RNAseq clusters. It creates an ROC curve for each cluster vs the others to help decide the appropriate number of clusters.
Installation
casc can be installed via devtools and github:
install.packages("devtools")
devtools::install_github("jamez-eh/casc")
Useage
casc has the following parameters:
sce: A SingleCellExperiment with normalized logcounts as an assay or a logcounts matrix with cells as columns and genes as rows.clusters: A list of clusterings to evaluate.alpha: A parameter for logistic regression. alpha = 1 represents the lasso penalty and alpha = 0 represents the ridge penalty.
registerDoSEQ()
sce_sim <- readRDS("~/sce_sim.rds")
casc_list <- casc(sce = sce_sim,
clusters = list(clustering_1, clustering_2, clustering_3, clustering_4, clustering_5),
alpha = 0.5)
Author
James Hopkins
jamez-eh/casc documentation built on June 12, 2019, 1:43 a.m.
casc
casc trains a logisitic regression model on provided single-cell RNAseq clusters. It creates an ROC curve for each cluster vs the others to help decide the appropriate number of clusters.

Installation
casc can be installed via devtools and github:
install.packages("devtools")
devtools::install_github("jamez-eh/casc")
Useage
casc has the following parameters:
sce: ASingleCellExperimentwith normalized logcounts as an assay or a logcounts matrix with cells as columns and genes as rows.clusters: A list of clusterings to evaluate.alpha: A parameter for logistic regression. alpha = 1 represents the lasso penalty and alpha = 0 represents the ridge penalty.
registerDoSEQ()
sce_sim <- readRDS("~/sce_sim.rds")
casc_list <- casc(sce = sce_sim,
clusters = list(clustering_1, clustering_2, clustering_3, clustering_4, clustering_5),
alpha = 0.5)
Author
James Hopkins
Embedding an R snippet on your website
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
