In joelcarlson/radiomics: 'Radiomic' Image Processing Toolbox
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>",
fig.path = "figs/README-"
)
Radiomics: Texture Analysis Matrices
Not Currently Maintained
This project is not currently being maintained. While I will do my best to help in a timely fashion, you should not expect a prompt response.
The radiomics package is a set of tools for computing texture matrices and features from images.
The release version of this package (April 2016, v0.1.2) is available from CRAN using:
install.packages("radiomics")
Or you can install the development version of the package using:
devtools::load_all(".")
devtools::install_github("joelcarlson/radiomics")
library(radiomics)
Texture Matrices
In the package are functions for calculating four different types of matrices and associated feature sets used to quantify the texture of an image.
These matrices are the:
- Grey Level Co-occurrence Matrix
- Grey Level Run Length Matrix
- Grey Level Size Zone Matrix
- Multiple Grey Level Size Zone Matrix
Detailed usage directions for calculating features and matrices can be found in the package vignette (use browseVignettes(package = "radiomics"))
Using the Package
Building Texture Matrices
Texture matrices can be created from 2D images by using the abbreviated and lowercase matrix name as a function call:
tumor <- radiomics::tumor #2D MRI slice of a brain tumor
glcm(tumor)
glrlm(tumor)
glszm(tumor)
mglszm(tumor)
A matrix with the class of the texture matrix type is returned, as shown here using glcm(tumor, n_grey=4)
glcm(tumor, n_grey=4)
class(glcm(tumor, n_grey=4))[1]
Visualizing Texture Matrices
Each matrix type has an associated image function for visualization of the results:
image(glcm(tumor))
image(glrlm(tumor))
image(glszm(tumor))
image(mglszm(tumor))
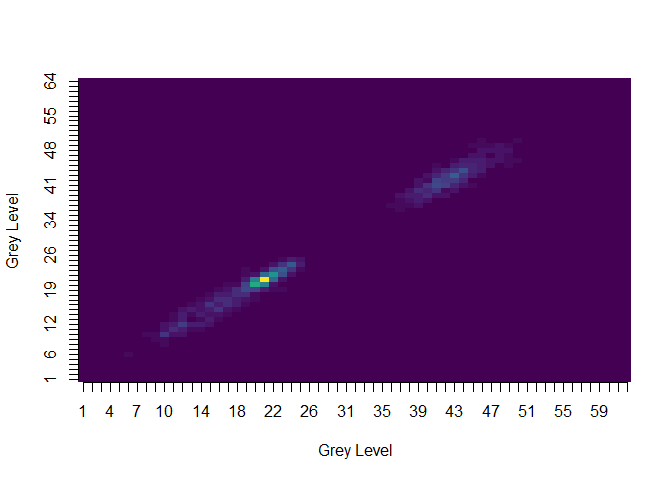
The image functions make use of the viridis scale, as shown here using image(glcm(tumor, n_grey=64)):
Calculating Features
Each matrix type has an associated calc_features function, which returns an object of class data.frame with a single observation for each calculated feature. First order features can also be calculated on 2D matrices.
calc_features(tumor)
calc_features(glcm(tumor))
calc_features(glrlm(tumor))
calc_features(glszm(tumor))
calc_features(mglszm(tumor))
joelcarlson/radiomics documentation built on May 19, 2019, 2:59 p.m.
knitr::opts_chunk$set( collapse = TRUE, comment = "#>", fig.path = "figs/README-" )
Radiomics: Texture Analysis Matrices
Not Currently Maintained
This project is not currently being maintained. While I will do my best to help in a timely fashion, you should not expect a prompt response.
The radiomics package is a set of tools for computing texture matrices and features from images.
The release version of this package (April 2016, v0.1.2) is available from CRAN using:
install.packages("radiomics")
Or you can install the development version of the package using:
devtools::load_all(".")
devtools::install_github("joelcarlson/radiomics") library(radiomics)
Texture Matrices
In the package are functions for calculating four different types of matrices and associated feature sets used to quantify the texture of an image.
These matrices are the:
- Grey Level Co-occurrence Matrix
- Grey Level Run Length Matrix
- Grey Level Size Zone Matrix
- Multiple Grey Level Size Zone Matrix
Detailed usage directions for calculating features and matrices can be found in the package vignette (use browseVignettes(package = "radiomics"))
Using the Package
Building Texture Matrices
Texture matrices can be created from 2D images by using the abbreviated and lowercase matrix name as a function call:
tumor <- radiomics::tumor #2D MRI slice of a brain tumor glcm(tumor) glrlm(tumor) glszm(tumor) mglszm(tumor)
A matrix with the class of the texture matrix type is returned, as shown here using glcm(tumor, n_grey=4)
glcm(tumor, n_grey=4)
class(glcm(tumor, n_grey=4))[1]
Visualizing Texture Matrices
Each matrix type has an associated image function for visualization of the results:
image(glcm(tumor)) image(glrlm(tumor)) image(glszm(tumor)) image(mglszm(tumor))
The image functions make use of the viridis scale, as shown here using image(glcm(tumor, n_grey=64)):

Calculating Features
Each matrix type has an associated calc_features function, which returns an object of class data.frame with a single observation for each calculated feature. First order features can also be calculated on 2D matrices.
calc_features(tumor) calc_features(glcm(tumor)) calc_features(glrlm(tumor)) calc_features(glszm(tumor)) calc_features(mglszm(tumor))
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
