README.md
In cmpsR: R Implementation of Congruent Matching Profile Segments Method
cmpsR 
The cmpsR package is an implementation of the Congruent Matching
Profile Segments (CMPS) method (Chen et al. 2019). In general, it can be
used for objective comparison of striated tool marks, but in our
examples, we mainly use it for bullet signatures comparison. The CMPS
score is expected to be large if two signatures are similar. So it can
also be considered as a feature that measures the similarity of two
bullet signatures.
Installation
You can install the released version of cmpsR from
CRAN with:
install.packages("cmpsR")
And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("willju-wangqian/cmpsR")
Example and Summary of the Algorithm
In this section we use a known match (KM) compasison (of two bullets) to
illustrate the main ideas of CMPS algorithm and to showcase the cmpsR
implementation. The cmpsR package includes a simple data set of 12
bullet signatures generated from two bullets (each bullet has 6 bullet
signatures). These bullet data come from the James Hamby Consecutively
Rifled Ruger Barrel Study (Brundage 1998; Hamby, Brundage, and Thorpe
2009; Hamby et al. 2019), and the two bullets included in cmpsR are
just a subset of Hamby set 252. These bullet data in their original
format can also be found in Chapter 3.5 of Open Forensic Science in
R.
A comparison of two bullets is considered as a match if two bullets are
fired from the same barrel (come from the same source). The gun barrel
used in the Hamby study has 6 lands, and during the firing process
striation marks will be engraved on the bullet by these lands. A bullet
signature is a numerical representation of the striation marks engraved
by a land. This is why each bullet can generate 6 bullet signatures. Two
bullet signatures are a match if they are originally engraved by the
same land in a gun barrel. Therefore, two bullets of a known-match
comparison will have 36 pairwise bullet signature comparisons, and 6 of
them are matching bullet signature comparisons while 30 of them are
non-matching bullet signature comparisons.
Here we plot the twelve bullet signatures of the two bullets. The bullet
signatures are aligned so that the top figure and the bottom figure of
the same column are a matching bullet signature comparison.

To further illustrate the idea of the CMPS algorithm, let’s consider one
matching bullet signature comparison: bullet signature of bullet 1 land
2 and bullet signature of bullet 2 land 3 (the second column), and
compute the CMPS score of this comparison:
library(cmpsR)
data("bullets")
x <- bullets$sigs[bullets$bulletland == "2-3"][[1]]$sig
y <- bullets$sigs[bullets$bulletland == "1-2"][[1]]$sig
cmps <- extract_feature_cmps(x, y, include = "full_result")
cmps$CMPS_score
#> [1] 18
And we have the plot of x and y.

Main Idea
The main idea of the CMPS method is that:
- we take the first signature as the comparison signature (
x or
bullet signature of “2-3”) and cut it into consecutive and
non-overlapping basis segments of the same length. In this case, we
set the length of a basis segment to be 50 units, and we have 22
basis segments in total for bullet signature x.

- for each basis segment, we compute the cross-correlation function
(ccf) between the basis segment and the reference signature (
y or
bullet signature of “1-2”)


- for the
ccf curve, the position represents the shift of the
segment. A negative value means a shift to the left, a positive
value means a shift to the right, and 0 means no shift (the segment
stays at its original position in the reference signature);
-
we are interested in the peaks in the ccf curve and the positions of
those peaks (as indicated by the red vertical line in the plot
above). In other words, if we shift the segment, which position
would give us the “best fit”?
-
If two signatures are from a KM comparison, most of the basis
segments should agree with each other on the position of the best
fit. Then these segments are called the “Congruent Matching
Profile Segments (CMPS)”.
Ideally, if two signatures are identical, we are expecting the position
of the highest peak in the ccf curve remains the same across all ccf
curves (we only show 7 segments here);

But in the real case, the basis segments might not achieve a final
agreement, but we have the majority;

We mark the 5 highest peaks for each ccf curve because the position of
the “highest peak” might not be the best one.
-
each ccf curve votes for 5 candidate positions, then we ask two
questions in order to obtain the CMPS number/score:
-
which position receives the most votes? -> the best position
(indicated by the red vertical line)
-
how many segments have voted for the best position? -> CMPS score
If we focus on these 7 segments only, and have a very short
tolerance zone, the CMPS number is 6.
(If we consider all 22 segments, and have a default tolerance zone
(+/- 25 units), the CMPS number is 20.)
-
false positive: how can the segments vote more wisely? -> Multi
Segment Lengths Strategy
-
by increasing the segment length, one can reduce the number of
“false positive” peaks.
-
the first scale level is the original length of segment 7; for the
second scale level, we double its length while keeping its center.
That is, we include 25 more units from both the left and right side
of the segment 7 to obtain a segment of 100 units length. For the
third scale level, we double the segment length again to obtain a
segment of length 200.


-
we choose five peaks at scale level 1; three peaks at scale level 2;
one peak at scale level 3
the peak shared by all three scale levels is a consistent
correlation peak (ccp). And the position of the ccp is our best
choice. Sometimes a ccp might not be found. Trying to identify a ccp
for each basis segment is called a “multi segment lengths” strategy.
The following plots (generated by cmpsR::cmps_segment_plot)
summarize the information of the two above plots. It shows that
segment 7 finds a consistent correlation peak (ccp) at a position
near 0 (position -6).
r
cmps <- extract_feature_cmps(x, y, include = "full_result")
cmps_plot_list <- cmpsR::cmps_segment_plot(cmps, seg_idx = 7)
ggpubr::ggarrange(plotlist = unlist(cmps_plot_list, recursive = FALSE),
nrow = 3, ncol = 2)

-
In this case, since segment 7 identifies a ccp, it casts a vote for
position -6. Then we ask two questions:
- which position receives the most votes (within a tolerance zone
specified by
Tx)?
- how many segments have voted for this position? -> CMPS score
-
by default, CMPS algorithm uses the multi-segment lengths strategy.
Use ?cmpsR::extract_feature_cmps to learn more about the function,
including its default settings.
-
If we follow the procedure described above (using the multi-segment
lengths strategy) and investigate all 22 basis segments, we can find
that 18 of them have cast a vote for position 0
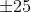 (since
(since Tx = 25 by default). Therefore, for this KM bullet
signature comparison, the CMPS score is 18.
cmps <- extract_feature_cmps(x, y, seg_length = 50, Tx = 25,
npeaks_set = c(5, 3, 1), include = "full_result")
cmps$CMPS_score
#> [1] 18
-
Segment 6 doesn’t cast a vote. Take a look at the following plot to
find out why.
r
cmps_plot_list <- cmpsR::cmps_segment_plot(cmps, seg_idx = 6)
ggpubr::ggarrange(plotlist = unlist(cmps_plot_list, recursive = FALSE),
nrow = 3, ncol = 2)

It doesn’t identify a consistent correlation peak.
-
If we have a KNM (known non-match) comparison, e.g. compare bullet
signature 2-3 with 1-3:
land23 <- bullets$sigs[bullets$bulletland == "2-3"][[1]]
land13 <- bullets$sigs[bullets$bulletland == "1-3"][[1]]
cmps_knm <- extract_feature_cmps(land23$sig, land13$sig, seg_length = 50, Tx = 25,
npeaks_set = c(5, 3, 1), include="full_result")
cmps_knm$CMPS_score
#> [1] 2
Full Comparison Between Two Bullets
extract_feature_cmps() can also be used in a pipeline fashion. The
following code performs a full comparison of two bullets. That is, it
evaluates all 36 pairwise bullet signature comparisons and computes the
CMPS scores.
library(tidyverse)
library(cmpsR)
data("bullets")
lands <- unique(bullets$bulletland)
comparisons <- data.frame(expand.grid(land1 = lands[1:6], land2 = lands[7:12]),
stringsAsFactors = FALSE)
comparisons <- comparisons %>%
left_join(bullets %>% select(bulletland, sig1=sigs),
by = c("land1" = "bulletland")) %>%
left_join(bullets %>% select(bulletland, sig2=sigs),
by = c("land2" = "bulletland"))
comparisons <- comparisons %>% mutate(
cmps = purrr::map2(sig1, sig2, .f = function(x, y) {
extract_feature_cmps(x$sig, y$sig, include = "full_result")
})
)
comparisons <- comparisons %>%
mutate(
cmps_score = sapply(comparisons$cmps, function(x) x$CMPS_score),
cmps_nseg = sapply(comparisons$cmps, function(x) x$nseg)
)
cp1 <- comparisons %>% select(land1, land2, cmps_score, cmps_nseg)
cp1
#> land1 land2 cmps_score cmps_nseg
#> 1 1-1 2-1 2 23
#> 2 1-2 2-1 2 22
#> 3 1-3 2-1 1 21
#> 4 1-4 2-1 2 22
#> 5 1-5 2-1 2 23
#> 6 1-6 2-1 16 22
#> 7 1-1 2-2 3 23
#> 8 1-2 2-2 1 22
#> 9 1-3 2-2 1 21
#> 10 1-4 2-2 1 22
#> 11 1-5 2-2 2 23
#> 12 1-6 2-2 3 22
#> 13 1-1 2-3 2 23
#> 14 1-2 2-3 17 22
#> 15 1-3 2-3 3 21
#> 16 1-4 2-3 1 22
#> 17 1-5 2-3 1 23
#> 18 1-6 2-3 1 22
#> 19 1-1 2-4 2 23
#> 20 1-2 2-4 1 22
#> 21 1-3 2-4 14 21
#> 22 1-4 2-4 1 22
#> 23 1-5 2-4 1 23
#> 24 1-6 2-4 2 22
#> 25 1-1 2-5 1 23
#> 26 1-2 2-5 2 22
#> 27 1-3 2-5 1 21
#> 28 1-4 2-5 10 22
#> 29 1-5 2-5 1 23
#> 30 1-6 2-5 1 22
#> 31 1-1 2-6 2 23
#> 32 1-2 2-6 3 22
#> 33 1-3 2-6 1 21
#> 34 1-4 2-6 1 22
#> 35 1-5 2-6 15 23
#> 36 1-6 2-6 1 22
The following plot summarizes the CMPS scores computed above.

Reference
Brundage, David J. 1998. “The Identification of
Consecutively Rifled Gun Barrels.” *AFTE Journal* 30 (3): 438–44.
Chen, Zhe, Wei Chu, Johannes A Soons, Robert M Thompson, John Song, and
Xuezeng Zhao. 2019. “Fired Bullet Signature
Correlation Using the Congruent Matching Profile Segments (CMPS)
Method.” *Forensic Science International*, December, \#109964.
.
Hamby, James E., David J. Brundage, Nicholas D. K. Petraco, and James W.
Thorpe. 2019. “A Worldwide Study of Bullets Fired From 10 Consecutively
Rifled 9mm RUGER Pistol Barrels—Analysis of Examiner Error Rate.”
*Journal of Forensic Sciences* 64 (2): 551–57.
.
Hamby, James E., David J. Brundage, and James W. Thorpe. 2009. “The Identification of Bullets Fired from 10 Consecutively
Rifled 9mm Ruger Pistol Barrels: A Research Project Involving 507
Participants from 20 Countries.” *AFTE Journal* 41 (2): 99–110.
Try the cmpsR package in your browser
Any scripts or data that you put into this service are public.
cmpsR documentation built on July 18, 2022, 9:07 a.m.
cmpsR 
The cmpsR package is an implementation of the Congruent Matching
Profile Segments (CMPS) method (Chen et al. 2019). In general, it can be
used for objective comparison of striated tool marks, but in our
examples, we mainly use it for bullet signatures comparison. The CMPS
score is expected to be large if two signatures are similar. So it can
also be considered as a feature that measures the similarity of two
bullet signatures.
Installation
You can install the released version of cmpsR from CRAN with:
install.packages("cmpsR")
And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("willju-wangqian/cmpsR")
Example and Summary of the Algorithm
In this section we use a known match (KM) compasison (of two bullets) to
illustrate the main ideas of CMPS algorithm and to showcase the cmpsR
implementation. The cmpsR package includes a simple data set of 12
bullet signatures generated from two bullets (each bullet has 6 bullet
signatures). These bullet data come from the James Hamby Consecutively
Rifled Ruger Barrel Study (Brundage 1998; Hamby, Brundage, and Thorpe
2009; Hamby et al. 2019), and the two bullets included in cmpsR are
just a subset of Hamby set 252. These bullet data in their original
format can also be found in Chapter 3.5 of Open Forensic Science in
R.
A comparison of two bullets is considered as a match if two bullets are fired from the same barrel (come from the same source). The gun barrel used in the Hamby study has 6 lands, and during the firing process striation marks will be engraved on the bullet by these lands. A bullet signature is a numerical representation of the striation marks engraved by a land. This is why each bullet can generate 6 bullet signatures. Two bullet signatures are a match if they are originally engraved by the same land in a gun barrel. Therefore, two bullets of a known-match comparison will have 36 pairwise bullet signature comparisons, and 6 of them are matching bullet signature comparisons while 30 of them are non-matching bullet signature comparisons.
Here we plot the twelve bullet signatures of the two bullets. The bullet signatures are aligned so that the top figure and the bottom figure of the same column are a matching bullet signature comparison.

To further illustrate the idea of the CMPS algorithm, let’s consider one matching bullet signature comparison: bullet signature of bullet 1 land 2 and bullet signature of bullet 2 land 3 (the second column), and compute the CMPS score of this comparison:
library(cmpsR)
data("bullets")
x <- bullets$sigs[bullets$bulletland == "2-3"][[1]]$sig
y <- bullets$sigs[bullets$bulletland == "1-2"][[1]]$sig
cmps <- extract_feature_cmps(x, y, include = "full_result")
cmps$CMPS_score
#> [1] 18
And we have the plot of x and y.

Main Idea
The main idea of the CMPS method is that:
- we take the first signature as the comparison signature (
xor bullet signature of “2-3”) and cut it into consecutive and non-overlapping basis segments of the same length. In this case, we set the length of a basis segment to be 50 units, and we have 22 basis segments in total for bullet signaturex.

- for each basis segment, we compute the cross-correlation function
(ccf) between the basis segment and the reference signature (
yor bullet signature of “1-2”)


- for the
ccfcurve, thepositionrepresents the shift of the segment. A negative value means a shift to the left, a positive value means a shift to the right, and 0 means no shift (the segment stays at its original position in the reference signature); -
we are interested in the peaks in the ccf curve and the positions of those peaks (as indicated by the red vertical line in the plot above). In other words, if we shift the segment, which position would give us the “best fit”?
-
If two signatures are from a KM comparison, most of the basis segments should agree with each other on the position of the best fit. Then these segments are called the “Congruent Matching Profile Segments (CMPS)”.
Ideally, if two signatures are identical, we are expecting the position of the highest peak in the ccf curve remains the same across all ccf curves (we only show 7 segments here);

But in the real case, the basis segments might not achieve a final agreement, but we have the majority;

We mark the 5 highest peaks for each ccf curve because the position of the “highest peak” might not be the best one.
-
each ccf curve votes for 5 candidate positions, then we ask two questions in order to obtain the CMPS number/score:
-
which position receives the most votes? -> the best position (indicated by the red vertical line)
-
how many segments have voted for the best position? -> CMPS score
If we focus on these 7 segments only, and have a very short tolerance zone, the CMPS number is 6.
(If we consider all 22 segments, and have a default tolerance zone (+/- 25 units), the CMPS number is 20.)
-
false positive: how can the segments vote more wisely? -> Multi Segment Lengths Strategy
-
by increasing the segment length, one can reduce the number of “false positive” peaks.
-
the first scale level is the original length of segment 7; for the second scale level, we double its length while keeping its center. That is, we include 25 more units from both the left and right side of the segment 7 to obtain a segment of 100 units length. For the third scale level, we double the segment length again to obtain a segment of length 200.


-
we choose five peaks at scale level 1; three peaks at scale level 2; one peak at scale level 3
the peak shared by all three scale levels is a consistent correlation peak (ccp). And the position of the ccp is our best choice. Sometimes a ccp might not be found. Trying to identify a ccp for each basis segment is called a “multi segment lengths” strategy.
The following plots (generated by
cmpsR::cmps_segment_plot) summarize the information of the two above plots. It shows that segment 7 finds a consistent correlation peak (ccp) at a position near 0 (position-6).r cmps <- extract_feature_cmps(x, y, include = "full_result") cmps_plot_list <- cmpsR::cmps_segment_plot(cmps, seg_idx = 7) ggpubr::ggarrange(plotlist = unlist(cmps_plot_list, recursive = FALSE), nrow = 3, ncol = 2)
-
In this case, since segment 7 identifies a ccp, it casts a vote for position
-6. Then we ask two questions:- which position receives the most votes (within a tolerance zone
specified by
Tx)? - how many segments have voted for this position? -> CMPS score
- which position receives the most votes (within a tolerance zone
specified by
-
by default, CMPS algorithm uses the multi-segment lengths strategy. Use
?cmpsR::extract_feature_cmpsto learn more about the function, including its default settings. -
If we follow the procedure described above (using the multi-segment lengths strategy) and investigate all 22 basis segments, we can find that 18 of them have cast a vote for position
0(since
Tx = 25by default). Therefore, for this KM bullet signature comparison, the CMPS score is 18.
cmps <- extract_feature_cmps(x, y, seg_length = 50, Tx = 25,
npeaks_set = c(5, 3, 1), include = "full_result")
cmps$CMPS_score
#> [1] 18
-
Segment 6 doesn’t cast a vote. Take a look at the following plot to find out why.
r cmps_plot_list <- cmpsR::cmps_segment_plot(cmps, seg_idx = 6) ggpubr::ggarrange(plotlist = unlist(cmps_plot_list, recursive = FALSE), nrow = 3, ncol = 2)
It doesn’t identify a consistent correlation peak.
-
If we have a KNM (known non-match) comparison, e.g. compare bullet signature 2-3 with 1-3:
land23 <- bullets$sigs[bullets$bulletland == "2-3"][[1]]
land13 <- bullets$sigs[bullets$bulletland == "1-3"][[1]]
cmps_knm <- extract_feature_cmps(land23$sig, land13$sig, seg_length = 50, Tx = 25,
npeaks_set = c(5, 3, 1), include="full_result")
cmps_knm$CMPS_score
#> [1] 2
Full Comparison Between Two Bullets
extract_feature_cmps() can also be used in a pipeline fashion. The
following code performs a full comparison of two bullets. That is, it
evaluates all 36 pairwise bullet signature comparisons and computes the
CMPS scores.
library(tidyverse)
library(cmpsR)
data("bullets")
lands <- unique(bullets$bulletland)
comparisons <- data.frame(expand.grid(land1 = lands[1:6], land2 = lands[7:12]),
stringsAsFactors = FALSE)
comparisons <- comparisons %>%
left_join(bullets %>% select(bulletland, sig1=sigs),
by = c("land1" = "bulletland")) %>%
left_join(bullets %>% select(bulletland, sig2=sigs),
by = c("land2" = "bulletland"))
comparisons <- comparisons %>% mutate(
cmps = purrr::map2(sig1, sig2, .f = function(x, y) {
extract_feature_cmps(x$sig, y$sig, include = "full_result")
})
)
comparisons <- comparisons %>%
mutate(
cmps_score = sapply(comparisons$cmps, function(x) x$CMPS_score),
cmps_nseg = sapply(comparisons$cmps, function(x) x$nseg)
)
cp1 <- comparisons %>% select(land1, land2, cmps_score, cmps_nseg)
cp1
#> land1 land2 cmps_score cmps_nseg
#> 1 1-1 2-1 2 23
#> 2 1-2 2-1 2 22
#> 3 1-3 2-1 1 21
#> 4 1-4 2-1 2 22
#> 5 1-5 2-1 2 23
#> 6 1-6 2-1 16 22
#> 7 1-1 2-2 3 23
#> 8 1-2 2-2 1 22
#> 9 1-3 2-2 1 21
#> 10 1-4 2-2 1 22
#> 11 1-5 2-2 2 23
#> 12 1-6 2-2 3 22
#> 13 1-1 2-3 2 23
#> 14 1-2 2-3 17 22
#> 15 1-3 2-3 3 21
#> 16 1-4 2-3 1 22
#> 17 1-5 2-3 1 23
#> 18 1-6 2-3 1 22
#> 19 1-1 2-4 2 23
#> 20 1-2 2-4 1 22
#> 21 1-3 2-4 14 21
#> 22 1-4 2-4 1 22
#> 23 1-5 2-4 1 23
#> 24 1-6 2-4 2 22
#> 25 1-1 2-5 1 23
#> 26 1-2 2-5 2 22
#> 27 1-3 2-5 1 21
#> 28 1-4 2-5 10 22
#> 29 1-5 2-5 1 23
#> 30 1-6 2-5 1 22
#> 31 1-1 2-6 2 23
#> 32 1-2 2-6 3 22
#> 33 1-3 2-6 1 21
#> 34 1-4 2-6 1 22
#> 35 1-5 2-6 15 23
#> 36 1-6 2-6 1 22
The following plot summarizes the CMPS scores computed above.

Reference
Try the cmpsR package in your browser
Any scripts or data that you put into this service are public.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
