In SchlossLab/mikropml: User-Friendly R Package for Supervised Machine Learning Pipelines
NOT_CRAN <- !identical(tolower(Sys.getenv("NOT_CRAN")), "false")
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>",
warning = FALSE,
purl = NOT_CRAN,
eval = NOT_CRAN
)
In this tutorial, we show how you can speed up pre-processing, model training,
and feature importance steps for individual runs, as well as how to train
multiple models in parallel within R and visualize the results.
However, we highly recommend using a workflow manager such as Snakemake rather
than parallelizing within a single R session.
Jump to the section Parallelizing with Snakemake
below if you're interested in skipping right to our best recommendation.
library(mikropml)
library(dplyr)
library(ggplot2)
Speed up single runs
By default, preprocess_data(), run_ml(), and compare_models() use only one process in series.
If you'd like to parallelize various steps of the pipeline to make them run
faster, install foreach, future, future.apply, and doFuture. Then,
register a future plan prior to calling these functions:
doFuture::registerDoFuture()
future::plan(future::multicore, workers = 2)
Above, we used the multicore plan to split the work across 2 cores. See the
future
documentation
for more about picking the best plan for your use case. Notably, multicore
does not work inside RStudio or on Windows; you will need to use multisession
instead in those cases.
After registering a future plan, you can call preprocess_data() and run_ml()
as usual, and they will run certain tasks in parallel.
otu_data_preproc <- preprocess_data(otu_mini_bin, "dx")$dat_transformed
result1 <- run_ml(otu_data_preproc, "glmnet", seed = 2019)
otu_data_preproc <- otu_data_preproc$dat_transformed
result1 <- otu_mini_bin_results_glmnet
There's a also a parallel version of the rf engine called parRF which trains
the trees in the forest in parallel. See the caret docs
for more information.
Bootstrap performance
If you only intend to call run_ml() once to generate one train/test split
(e.g. such as for a temporal split of the dataset), you can evaluate the
model performance by bootstrapping the test set.
Here we show how to generate 100 bootstraps and calculate a confidence
interval for the model performance. We only use 100 here for computation
speed, but it is recommended to generate 10000 bootstraps for a more precise
estimation of the confidence interval.
boot_perf <- bootstrap_performance(result1,
outcome_colname = "dx",
bootstrap_times = 100, alpha = 0.05
)
boot_perf
Call run_ml() multiple times in parallel in R
You can use functions from the future.apply package to call run_ml()
multiple times in parallel with different parameters. You will first need to run
future::plan() as above if you haven't already. Then, call run_ml() with
multiple seeds using future_lapply():
# NOTE: use more seeds for real-world data
results_multi <- future.apply::future_lapply(seq(100, 102), function(seed) {
run_ml(otu_data_preproc, "glmnet", seed = seed)
}, future.seed = TRUE)
Each call to run_ml() with a different seed uses a different random split of
the data into training and testing sets. Since we are using seeds, we must set
future.seed to TRUE (see the future.apply
documentation
and this blog
post
for details on parallel-safe random seeds). This example uses only a few seeds
for speed and simplicity, but for real data we recommend using many more seeds
to get a better estimate of model performance.
In these examples, we used functions from the future.apply package to
run_ml() in parallel, but you can accomplish the same thing with parallel
versions of the purrr::map() functions using the furrr package (e.g.
furrr::future_map_dfr()).
Extract the performance results and combine into one dataframe for all seeds:
perf_df <- future.apply::future_lapply(results_multi,
function(result) {
result[["performance"]] %>%
select(cv_metric_AUC, AUC, method)
},
future.seed = TRUE
) %>%
dplyr::bind_rows()
perf_df
Multiple ML methods
You may also wish to compare performance for different ML methods. mapply()
can iterate over multiple lists or vectors, and future_mapply() works the same
way:
# NOTE: use more seeds for real-world data
param_grid <- expand.grid(
seeds = seq(100, 103),
methods = c("glmnet", "rf")
)
results_mtx <- future.apply::future_mapply(
function(seed, method) {
run_ml(otu_data_preproc,
method,
seed = seed,
find_feature_importance = TRUE
)
},
param_grid$seeds,
param_grid$methods %>% as.character(),
future.seed = TRUE
)
Visualize the results
ggplot2 is required to use our plotting functions below.
You can also create your own plots however you like using the results data.
Performance
Mean AUC
perf_df <- lapply(
results_mtx["performance", ],
function(x) {
x %>% select(cv_metric_AUC, AUC, method)
}
) %>%
dplyr::bind_rows()
perf_boxplot <- plot_model_performance(perf_df)
perf_boxplot
plot_model_performance() returns a ggplot2 object.
You can add layers to customize the plot:
perf_boxplot +
theme_classic() +
scale_color_brewer(palette = "Dark2") +
coord_flip()
ROC and PRC curves
First calculate the sensitivity, specificity, and precision for all models.
get_sensspec_seed <- function(colnum) {
result <- results_mtx[, colnum]
trained_model <- result$trained_model
test_data <- result$test_data
seed <- result$performance$seed
method <- result$trained_model$method
sensspec <- calc_model_sensspec(
trained_model,
test_data,
"dx"
) %>%
mutate(seed = seed, method = method)
return(sensspec)
}
sensspec_dat <- purrr::map_dfr(
seq(1, dim(results_mtx)[2]),
get_sensspec_seed
)
Plot curves for a single model
sensspec_1 <- sensspec_dat %>% filter(seed == 100, method == "glmnet")
sensspec_1 %>%
ggplot(aes(x = specificity, y = sensitivity, )) +
geom_line() +
geom_abline(
intercept = 1, slope = 1,
linetype = "dashed", color = "grey50"
) +
coord_equal() +
scale_x_reverse(expand = c(0, 0), limits = c(1.01, -0.01)) +
scale_y_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) +
labs(x = "Specificity", y = "Sensitivity") +
theme_bw() +
theme(legend.title = element_blank())
baseline_precision_otu <- calc_baseline_precision(
otu_data_preproc,
"dx", "cancer"
)
sensspec_1 %>%
rename(recall = sensitivity) %>%
ggplot(aes(x = recall, y = precision, )) +
geom_line() +
geom_hline(
yintercept = baseline_precision_otu,
linetype = "dashed", color = "grey50"
) +
coord_equal() +
scale_x_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) +
scale_y_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) +
labs(x = "Recall", y = "Precision") +
theme_bw() +
theme(legend.title = element_blank())
Plot mean ROC and PRC for all models
sensspec_dat %>%
calc_mean_roc() %>%
plot_mean_roc()
sensspec_dat %>%
calc_mean_prc() %>%
plot_mean_prc(baseline_precision = baseline_precision_otu)
Feature importance
The perf_metric_diff from the feature importance data frame contains the
differences between the performance on the actual test data and the performance
on the permuted test data (i.e. test minus permuted).
If a feature is important for model performance, we expect perf_metric_diff to
be positive.
In other words, the features that resulted in the largest decrease in
performance when permuted are the most important features.
Feature importance for multiple models
You can select the top n most important features for your models and plot them
like so:
feat_df <- results_mtx["feature_importance", ] %>%
dplyr::bind_rows()
top_n <- 5
top_feats <- feat_df %>%
group_by(method, feat) %>%
summarize(mean_diff = median(perf_metric_diff)) %>%
filter(mean_diff > 0) %>%
slice_max(order_by = mean_diff, n = top_n)
feat_df %>%
right_join(top_feats, by = c("method", "feat")) %>%
mutate(features = forcats::fct_reorder(factor(feat), mean_diff)) %>%
ggplot(aes(x = perf_metric_diff, y = features, color = method)) +
geom_boxplot() +
geom_vline(xintercept = 0, linetype = "dashed") +
labs(
x = "Decrease in performance (actual minus permutation)",
y = "Features",
caption = "Features which have a lower performance when permuted have a
difference in performance above zero. The features with the greatest
decrease are the most important for model performance." %>%
stringr::str_wrap(width = 100)
) +
theme_bw() +
theme(plot.caption = element_text(hjust = 0))
See the docs for get_feature_importance() for more details on how these values
are computed.
Feature importance for a single model
You can also plot feature importance for a single model.
Here we report the actual performance, the permutation performance, and the
empirical 95% confidence interval for the permutation performance.
feat_imp_1 <- results_mtx[, 1][["feature_importance"]]
perf_metric_name <- results_mtx[, 1][["trained_model"]]$metric
perf_actual <- results_mtx[, 1][["performance"]] %>% pull(perf_metric_name)
feat_imp_1 %>%
filter(perf_metric_diff > 0) %>%
mutate(feat = if_else(pvalue < 0.05, paste0("*", feat), as.character(feat)) %>%
as.factor() %>%
forcats::fct_reorder(perf_metric_diff)) %>%
ggplot(aes(x = perf_metric, xmin = lower, xmax = upper, y = feat)) +
geom_pointrange() +
geom_vline(xintercept = perf_actual, linetype = "dashed") +
labs(
x = "Permutation performance",
y = "Features",
caption = "The dashed line represents the actual performance on the
test set. Features which have a lower performance when permuted are
important for model performance. Significant features (pvalue < 0.05)
are marked with an asterisk (*). Error bars represent the 95%
confidence interval." %>% stringr::str_wrap(width = 110)
) +
theme_bw() +
theme(plot.caption = element_text(hjust = 0))
Live progress updates
preprocess_data() and get_feature_importance() support reporting live
progress updates using the progressr package. The format is up to you, but we
recommend using a progress bar like this:
# optionally, specify the progress bar format with the `progress` package.
progressr::handlers(progressr::handler_progress(
format = ":message :bar :percent | elapsed: :elapsed | eta: :eta",
clear = FALSE,
show_after = 0
))
# tell progressr to always report progress in any functions that use it.
# set this to FALSE to turn it back off again.
progressr::handlers(global = TRUE)
# run your code and watch the live progress updates.
dat <- preprocess_data(otu_mini_bin, "dx")$dat_transformed
#> Using 'dx' as the outcome column.
#> preprocessing ========================>------- 78% | elapsed: 1s | eta: 0s
results <- run_ml(dat, "glmnet",
kfold = 2, cv_times = 2,
find_feature_importance = TRUE
)
#> Using 'dx' as the outcome column.
#> Training the model...
#> Training complete.
#> Feature importance =========================== 100% | elapsed: 37s | eta: 0s
Note that some future backends support "near-live" progress updates, meaning the
progress may not be reported immediately when parallel processing with futures.
Read more on that in the progressr
vignette.
For more on progressr and how to customize the format of progress updates, see
the progressr docs.
Parallelizing with Snakemake
When parallelizing multiple calls to run_ml() in R as in the examples above,
all of the results objects are held in memory. This isn't a big deal for a small
dataset run with only a few seeds. However, for large datasets run in parallel
with, say, 100 seeds (recommended), you may run into problems trying to store
all of those objects in memory at once.
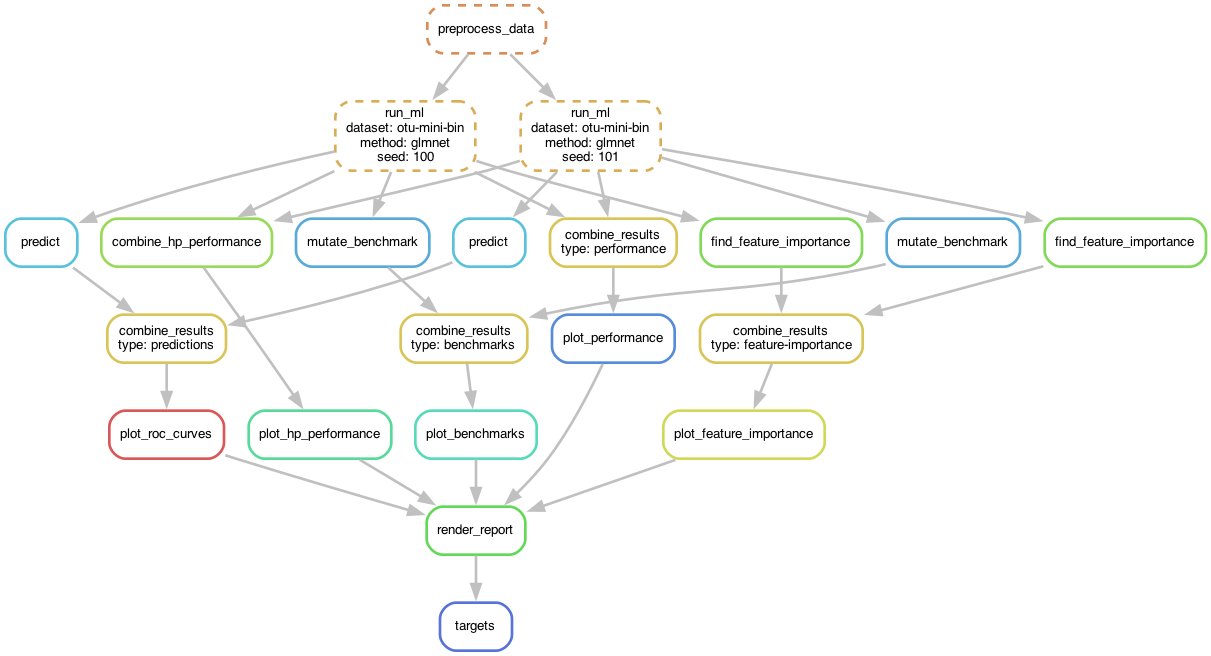
Using a workflow manager such as Snakemake or Nextflow is highly recommend to
maximize the scalability and reproducibility of computational analyses.
We created a template Snakemake workflow here
which you can use as a starting point for your ML project.
SchlossLab/mikropml documentation built on Aug. 24, 2023, 9:51 p.m.
NOT_CRAN <- !identical(tolower(Sys.getenv("NOT_CRAN")), "false") knitr::opts_chunk$set( collapse = TRUE, comment = "#>", warning = FALSE, purl = NOT_CRAN, eval = NOT_CRAN )
In this tutorial, we show how you can speed up pre-processing, model training, and feature importance steps for individual runs, as well as how to train multiple models in parallel within R and visualize the results. However, we highly recommend using a workflow manager such as Snakemake rather than parallelizing within a single R session. Jump to the section Parallelizing with Snakemake below if you're interested in skipping right to our best recommendation.
library(mikropml) library(dplyr) library(ggplot2)
Speed up single runs
By default, preprocess_data(), run_ml(), and compare_models() use only one process in series.
If you'd like to parallelize various steps of the pipeline to make them run
faster, install foreach, future, future.apply, and doFuture. Then,
register a future plan prior to calling these functions:
doFuture::registerDoFuture() future::plan(future::multicore, workers = 2)
Above, we used the multicore plan to split the work across 2 cores. See the
future
documentation
for more about picking the best plan for your use case. Notably, multicore
does not work inside RStudio or on Windows; you will need to use multisession
instead in those cases.
After registering a future plan, you can call preprocess_data() and run_ml()
as usual, and they will run certain tasks in parallel.
otu_data_preproc <- preprocess_data(otu_mini_bin, "dx")$dat_transformed result1 <- run_ml(otu_data_preproc, "glmnet", seed = 2019)
otu_data_preproc <- otu_data_preproc$dat_transformed result1 <- otu_mini_bin_results_glmnet
There's a also a parallel version of the rf engine called parRF which trains
the trees in the forest in parallel. See the caret docs
for more information.
Bootstrap performance
If you only intend to call run_ml() once to generate one train/test split
(e.g. such as for a temporal split of the dataset), you can evaluate the
model performance by bootstrapping the test set.
Here we show how to generate 100 bootstraps and calculate a confidence
interval for the model performance. We only use 100 here for computation
speed, but it is recommended to generate 10000 bootstraps for a more precise
estimation of the confidence interval.
boot_perf <- bootstrap_performance(result1, outcome_colname = "dx", bootstrap_times = 100, alpha = 0.05 ) boot_perf
Call run_ml() multiple times in parallel in R
You can use functions from the future.apply package to call run_ml()
multiple times in parallel with different parameters. You will first need to run
future::plan() as above if you haven't already. Then, call run_ml() with
multiple seeds using future_lapply():
# NOTE: use more seeds for real-world data results_multi <- future.apply::future_lapply(seq(100, 102), function(seed) { run_ml(otu_data_preproc, "glmnet", seed = seed) }, future.seed = TRUE)
Each call to run_ml() with a different seed uses a different random split of
the data into training and testing sets. Since we are using seeds, we must set
future.seed to TRUE (see the future.apply
documentation
and this blog
post
for details on parallel-safe random seeds). This example uses only a few seeds
for speed and simplicity, but for real data we recommend using many more seeds
to get a better estimate of model performance.
In these examples, we used functions from the future.apply package to
run_ml() in parallel, but you can accomplish the same thing with parallel
versions of the purrr::map() functions using the furrr package (e.g.
furrr::future_map_dfr()).
Extract the performance results and combine into one dataframe for all seeds:
perf_df <- future.apply::future_lapply(results_multi, function(result) { result[["performance"]] %>% select(cv_metric_AUC, AUC, method) }, future.seed = TRUE ) %>% dplyr::bind_rows() perf_df
Multiple ML methods
You may also wish to compare performance for different ML methods. mapply()
can iterate over multiple lists or vectors, and future_mapply() works the same
way:
# NOTE: use more seeds for real-world data param_grid <- expand.grid( seeds = seq(100, 103), methods = c("glmnet", "rf") ) results_mtx <- future.apply::future_mapply( function(seed, method) { run_ml(otu_data_preproc, method, seed = seed, find_feature_importance = TRUE ) }, param_grid$seeds, param_grid$methods %>% as.character(), future.seed = TRUE )
Visualize the results
ggplot2 is required to use our plotting functions below.
You can also create your own plots however you like using the results data.
Performance
Mean AUC
perf_df <- lapply( results_mtx["performance", ], function(x) { x %>% select(cv_metric_AUC, AUC, method) } ) %>% dplyr::bind_rows() perf_boxplot <- plot_model_performance(perf_df) perf_boxplot
plot_model_performance() returns a ggplot2 object.
You can add layers to customize the plot:
perf_boxplot + theme_classic() + scale_color_brewer(palette = "Dark2") + coord_flip()
ROC and PRC curves
First calculate the sensitivity, specificity, and precision for all models.
get_sensspec_seed <- function(colnum) { result <- results_mtx[, colnum] trained_model <- result$trained_model test_data <- result$test_data seed <- result$performance$seed method <- result$trained_model$method sensspec <- calc_model_sensspec( trained_model, test_data, "dx" ) %>% mutate(seed = seed, method = method) return(sensspec) } sensspec_dat <- purrr::map_dfr( seq(1, dim(results_mtx)[2]), get_sensspec_seed )
Plot curves for a single model
sensspec_1 <- sensspec_dat %>% filter(seed == 100, method == "glmnet") sensspec_1 %>% ggplot(aes(x = specificity, y = sensitivity, )) + geom_line() + geom_abline( intercept = 1, slope = 1, linetype = "dashed", color = "grey50" ) + coord_equal() + scale_x_reverse(expand = c(0, 0), limits = c(1.01, -0.01)) + scale_y_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) + labs(x = "Specificity", y = "Sensitivity") + theme_bw() + theme(legend.title = element_blank()) baseline_precision_otu <- calc_baseline_precision( otu_data_preproc, "dx", "cancer" ) sensspec_1 %>% rename(recall = sensitivity) %>% ggplot(aes(x = recall, y = precision, )) + geom_line() + geom_hline( yintercept = baseline_precision_otu, linetype = "dashed", color = "grey50" ) + coord_equal() + scale_x_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) + scale_y_continuous(expand = c(0, 0), limits = c(-0.01, 1.01)) + labs(x = "Recall", y = "Precision") + theme_bw() + theme(legend.title = element_blank())
Plot mean ROC and PRC for all models
sensspec_dat %>% calc_mean_roc() %>% plot_mean_roc() sensspec_dat %>% calc_mean_prc() %>% plot_mean_prc(baseline_precision = baseline_precision_otu)
Feature importance
The perf_metric_diff from the feature importance data frame contains the
differences between the performance on the actual test data and the performance
on the permuted test data (i.e. test minus permuted).
If a feature is important for model performance, we expect perf_metric_diff to
be positive.
In other words, the features that resulted in the largest decrease in
performance when permuted are the most important features.
Feature importance for multiple models
You can select the top n most important features for your models and plot them like so:
feat_df <- results_mtx["feature_importance", ] %>% dplyr::bind_rows() top_n <- 5 top_feats <- feat_df %>% group_by(method, feat) %>% summarize(mean_diff = median(perf_metric_diff)) %>% filter(mean_diff > 0) %>% slice_max(order_by = mean_diff, n = top_n) feat_df %>% right_join(top_feats, by = c("method", "feat")) %>% mutate(features = forcats::fct_reorder(factor(feat), mean_diff)) %>% ggplot(aes(x = perf_metric_diff, y = features, color = method)) + geom_boxplot() + geom_vline(xintercept = 0, linetype = "dashed") + labs( x = "Decrease in performance (actual minus permutation)", y = "Features", caption = "Features which have a lower performance when permuted have a difference in performance above zero. The features with the greatest decrease are the most important for model performance." %>% stringr::str_wrap(width = 100) ) + theme_bw() + theme(plot.caption = element_text(hjust = 0))
See the docs for get_feature_importance() for more details on how these values
are computed.
Feature importance for a single model
You can also plot feature importance for a single model. Here we report the actual performance, the permutation performance, and the empirical 95% confidence interval for the permutation performance.
feat_imp_1 <- results_mtx[, 1][["feature_importance"]] perf_metric_name <- results_mtx[, 1][["trained_model"]]$metric perf_actual <- results_mtx[, 1][["performance"]] %>% pull(perf_metric_name) feat_imp_1 %>% filter(perf_metric_diff > 0) %>% mutate(feat = if_else(pvalue < 0.05, paste0("*", feat), as.character(feat)) %>% as.factor() %>% forcats::fct_reorder(perf_metric_diff)) %>% ggplot(aes(x = perf_metric, xmin = lower, xmax = upper, y = feat)) + geom_pointrange() + geom_vline(xintercept = perf_actual, linetype = "dashed") + labs( x = "Permutation performance", y = "Features", caption = "The dashed line represents the actual performance on the test set. Features which have a lower performance when permuted are important for model performance. Significant features (pvalue < 0.05) are marked with an asterisk (*). Error bars represent the 95% confidence interval." %>% stringr::str_wrap(width = 110) ) + theme_bw() + theme(plot.caption = element_text(hjust = 0))
Live progress updates
preprocess_data() and get_feature_importance() support reporting live
progress updates using the progressr package. The format is up to you, but we
recommend using a progress bar like this:
# optionally, specify the progress bar format with the `progress` package. progressr::handlers(progressr::handler_progress( format = ":message :bar :percent | elapsed: :elapsed | eta: :eta", clear = FALSE, show_after = 0 )) # tell progressr to always report progress in any functions that use it. # set this to FALSE to turn it back off again. progressr::handlers(global = TRUE) # run your code and watch the live progress updates. dat <- preprocess_data(otu_mini_bin, "dx")$dat_transformed #> Using 'dx' as the outcome column. #> preprocessing ========================>------- 78% | elapsed: 1s | eta: 0s results <- run_ml(dat, "glmnet", kfold = 2, cv_times = 2, find_feature_importance = TRUE ) #> Using 'dx' as the outcome column. #> Training the model... #> Training complete. #> Feature importance =========================== 100% | elapsed: 37s | eta: 0s
Note that some future backends support "near-live" progress updates, meaning the
progress may not be reported immediately when parallel processing with futures.
Read more on that in the progressr
vignette.
For more on progressr and how to customize the format of progress updates, see
the progressr docs.
Parallelizing with Snakemake
When parallelizing multiple calls to run_ml() in R as in the examples above,
all of the results objects are held in memory. This isn't a big deal for a small
dataset run with only a few seeds. However, for large datasets run in parallel
with, say, 100 seeds (recommended), you may run into problems trying to store
all of those objects in memory at once.
Using a workflow manager such as Snakemake or Nextflow is highly recommend to maximize the scalability and reproducibility of computational analyses. We created a template Snakemake workflow here which you can use as a starting point for your ML project.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.