inst/examples/appendices/delayed.md
In cboettig/earlywarning: Detection of Early Warning Signals for Catastrophic Bifurcations
Model-based detection of early warning under a delay
What happens when we attempt to fit the linear change model to a signal in which the environment is initially constant and then begins degrading?
First let's simulate some such data
require(populationdynamics)
## Loading required package: populationdynamics
require(earlywarning)
## Loading required package: earlywarning
pars = c(Xo = 500, e = 0.5, a = 180, K = 1000,
h = 200, i = 0, Da = 0.45, Dt = 50, p = 2)
sn <- saddle_node_ibm(pars, times = seq(0, 100,
length = 200))
X <- ts(sn$x1, start = sn$time[1], deltat = sn$time[2] -
sn$time[1])
Observe that this produces timeseries has 200 points in the interval (0,100) with a linear change begining half way through, at Dt=50, where it begins approaching a saddle-node bifurcation. Let's fit both our models:
A <- stability_model(X, "OU")
B <- stability_model(X, "LSN")
observed <- -2 * (logLik(A) - logLik(B))
observed
## [1] 4.805
... and then use the bootstrapped likelihood ratios to see if this difference is significant:
require(snowfall)
## Loading required package: snowfall
## Loading required package: snow
sfInit(parallel = TRUE, cpu = 16)
## R Version: R version 2.14.1 (2011-12-22)
##
## snowfall 1.84 initialized (using snow 0.3-8): parallel execution on 16 CPUs.
##
sfLibrary(earlywarning)
## Library earlywarning loaded.
## Library earlywarning loaded in cluster.
##
## Warning message: 'keep.source' is deprecated and will be ignored
sfExportAll()
reps <- sfLapply(1:100, function(i) compare(A,
B))
lr <- lik_ratios(reps)
roc <- roc_data(lr)
save(list = ls(), file = "delayed.rda")
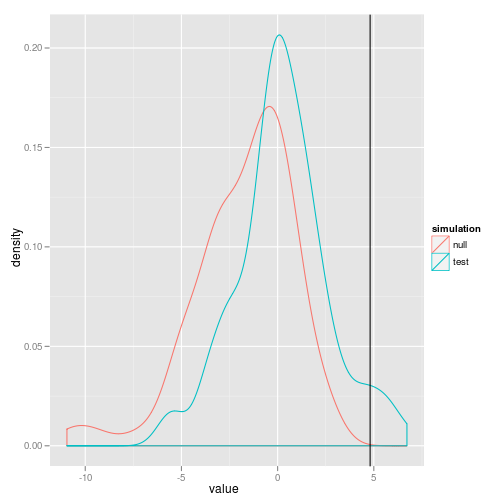
Plot the likelihood ratio distribution with a line indicating the observed value. Also plot the ROC curve.
require(ggplot2)
## Loading required package: ggplot2
## Loading required package: reshape
## Loading required package: plyr
##
## Attaching package: 'reshape'
##
## The following object(s) are masked from 'package:plyr':
##
## rename, round_any
##
## Loading required package: grid
## Loading required package: proto
ggplot(lr) + geom_density(aes(value, color = simulation)) +
geom_vline(aes(xintercept = observed))
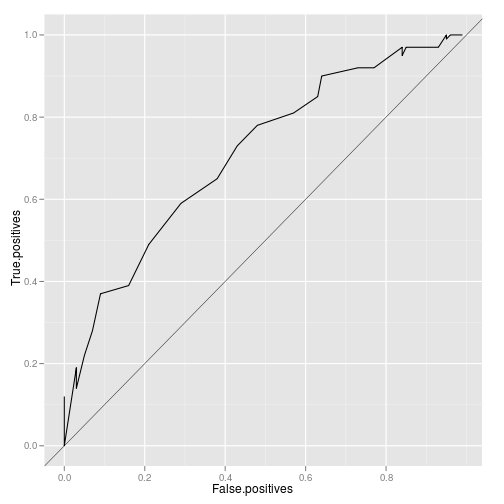
ggplot(roc) + geom_line(aes(False.positives, True.positives)) +
geom_abline(aes(yintercept = 0, slope = 1), lwd = 0.2)
Is this a concern for the windowed approach? It still complicates the calculation of the correlation statistic used to demonstrate a statistically significant increase. We certainly would not advocate choosing the region "by eye" as it were where the increase appears and then testing the correlation on that segment alone -- that would guarentee bias. In principle this faces the same problem that the starting point needs to be known in advance.
cboettig/earlywarning documentation built on May 13, 2019, 2:07 p.m.
Model-based detection of early warning under a delay
What happens when we attempt to fit the linear change model to a signal in which the environment is initially constant and then begins degrading?
First let's simulate some such data
require(populationdynamics)
## Loading required package: populationdynamics
require(earlywarning)
## Loading required package: earlywarning
pars = c(Xo = 500, e = 0.5, a = 180, K = 1000,
h = 200, i = 0, Da = 0.45, Dt = 50, p = 2)
sn <- saddle_node_ibm(pars, times = seq(0, 100,
length = 200))
X <- ts(sn$x1, start = sn$time[1], deltat = sn$time[2] -
sn$time[1])
Observe that this produces timeseries has 200 points in the interval (0,100) with a linear change begining half way through, at Dt=50, where it begins approaching a saddle-node bifurcation. Let's fit both our models:
A <- stability_model(X, "OU")
B <- stability_model(X, "LSN")
observed <- -2 * (logLik(A) - logLik(B))
observed
## [1] 4.805
... and then use the bootstrapped likelihood ratios to see if this difference is significant:
require(snowfall)
## Loading required package: snowfall
## Loading required package: snow
sfInit(parallel = TRUE, cpu = 16)
## R Version: R version 2.14.1 (2011-12-22)
##
## snowfall 1.84 initialized (using snow 0.3-8): parallel execution on 16 CPUs.
##
sfLibrary(earlywarning)
## Library earlywarning loaded.
## Library earlywarning loaded in cluster.
##
## Warning message: 'keep.source' is deprecated and will be ignored
sfExportAll()
reps <- sfLapply(1:100, function(i) compare(A,
B))
lr <- lik_ratios(reps)
roc <- roc_data(lr)
save(list = ls(), file = "delayed.rda")
Plot the likelihood ratio distribution with a line indicating the observed value. Also plot the ROC curve.
require(ggplot2)
## Loading required package: ggplot2
## Loading required package: reshape
## Loading required package: plyr
##
## Attaching package: 'reshape'
##
## The following object(s) are masked from 'package:plyr':
##
## rename, round_any
##
## Loading required package: grid
## Loading required package: proto
ggplot(lr) + geom_density(aes(value, color = simulation)) +
geom_vline(aes(xintercept = observed))

ggplot(roc) + geom_line(aes(False.positives, True.positives)) +
geom_abline(aes(yintercept = 0, slope = 1), lwd = 0.2)

Is this a concern for the windowed approach? It still complicates the calculation of the correlation statistic used to demonstrate a statistically significant increase. We certainly would not advocate choosing the region "by eye" as it were where the increase appears and then testing the correlation on that segment alone -- that would guarentee bias. In principle this faces the same problem that the starting point needs to be known in advance.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
