inst/examples/BUGS/process-noise-only.md
In cboettig/nonparametric-bayes: Nonparametric Bayes inference for ecological models
Comparison of Nonparametric Bayesian Gaussian Process estimates to standard the Parametric Bayesian approach
Plotting and knitr options, (can generally be ignored)
require(modeest)
posterior.mode <- function(x) {
mlv(x, method="shorth")$M
}
Model and parameters
Uses the model derived in citet("10.1080/10236190412331335373"), of a Ricker-like growth curve with an allee effect, defined in the pdgControl package,
f <- RickerAllee
p <- c(2, 8, 5)
K <- 10 # approx, a li'l' less
allee <- 5 # approx, a li'l' less
Various parameters defining noise dynamics, grid, and policy costs.
sigma_g <- 0.05
sigma_m <- 0.0
z_g <- function() rlnorm(1, 0, sigma_g)
z_m <- function() 1
x_grid <- seq(0, 1.5 * K, length=50)
h_grid <- x_grid
profit <- function(x,h) pmin(x, h)
delta <- 0.01
OptTime <- 50 # stationarity with unstable models is tricky thing
reward <- 0
xT <- 0
Xo <- allee+.5# observations start from
x0 <- K # simulation under policy starts from
Tobs <- 40
MaxT = 1000 # timeout for value iteration convergence
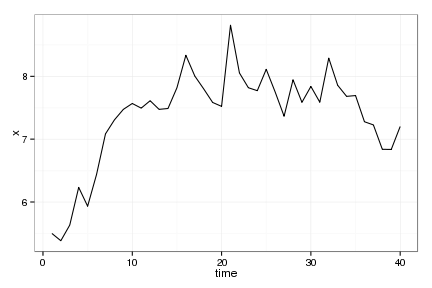
Sample Data
set.seed(1234)
#harvest <- sort(rep(seq(0, .5, length=7), 5))
x <- numeric(Tobs)
x[1] <- Xo
nz <- 1
for(t in 1:(Tobs-1))
x[t+1] = z_g() * f(x[t], h=0, p=p)
obs <- data.frame(x = c(rep(0,nz),
pmax(rep(0,Tobs-1), x[1:(Tobs-1)])),
y = c(rep(0,nz),
x[2:Tobs]))
raw_plot <- ggplot(data.frame(time = 1:Tobs, x=x), aes(time,x)) + geom_line()
raw_plot

Maximum Likelihood
set.seed(12345)
estf <- function(p){
mu <- f(obs$x,0,p)
-sum(dlnorm(obs$y, log(mu), p[4]), log=TRUE)
}
par <- c(p[1]*rlnorm(1,0,.1),
p[2]*rlnorm(1,0,.1),
p[3]*rlnorm(1,0, .1),
sigma_g * rlnorm(1,0,.1))
o <- optim(par, estf, method="L", lower=c(1e-5,1e-5,1e-5,1e-5))
f_alt <- f
p_alt <- c(as.numeric(o$par[1]), as.numeric(o$par[2]), as.numeric(o$par[3]))
sigma_g_alt <- as.numeric(o$par[4])
est <- list(f = f_alt, p = p_alt, sigma_g = sigma_g_alt, mloglik=o$value)
Mean predictions
true_means <- sapply(x_grid, f, 0, p)
est_means <- sapply(x_grid, est$f, 0, est$p)
Non-parametric Bayes
#inv gamma has mean b / (a - 1) (assuming a>1) and variance b ^ 2 / ((a - 2) * (a - 1) ^ 2) (assuming a>2)
s2.p <- c(5,5)
d.p = c(10, 1/0.1)
Estimate the Gaussian Process (nonparametric Bayesian fit)
gp <- gp_mcmc(obs$x, y=obs$y, n=1e5, s2.p = s2.p, d.p = d.p)
gp_dat <- gp_predict(gp, x_grid, burnin=1e4, thin=300)
Show traces and posteriors against priors
gp_assessment_plots <- summary_gp_mcmc(gp, burnin=1e4, thin=300)
# Summarize the GP model
tgp_dat <-
data.frame( x = x_grid,
y = gp_dat$E_Ef,
ymin = gp_dat$E_Ef - 2 * sqrt(gp_dat$E_Vf),
ymax = gp_dat$E_Ef + 2 * sqrt(gp_dat$E_Vf) )
Parametric Bayesian Models
We use the JAGS Gibbs sampler, a recent open source BUGS
implementation with an R interface that works on most platforms.
We initialize the usual MCMC parameters; see ?jags for details.
All parametric Bayesian estimates use the following basic parameters for the JAGS MCMC:
y <- x
N <- length(x);
jags.data <- list("N","y")
n.chains <- 6
n.iter <- 1e6
n.burnin <- floor(10000)
n.thin <- max(1, floor(n.chains * (n.iter - n.burnin)/1000))
n.update <- 10
We will use the same priors for process and observation noise in each model,
stdQ_prior_p <- c(1e-6, 100)
stdR_prior_p <- c(1e-6, .1)
stdQ_prior <- function(x) dunif(x, stdQ_prior_p[1], stdQ_prior_p[2])
stdR_prior <- function(x) dunif(x, stdR_prior_p[1], stdR_prior_p[2])
Parametric Bayes of correct (Allen) model
We initiate the MCMC chain (init_p) using the true values of the
parameters p from the simulation. While impossible in real data, this
gives the parametric Bayesian approach the best chance at succeeding.
y is the timeseries (recall obs has the $x_t$, $x_{t+1}$ pairs)
The actual model is defined in a model.file that contains an R function
that is automatically translated into BUGS code by R2WinBUGS. The file
defines the priors and the model. We write the file from R as follows:
K_prior_p <- c(0.01, 20.0)
r0_prior_p <- c(0.01, 6.0)
theta_prior_p <- c(0.01, 20.0)
bugs.model <-
paste(sprintf(
"model{
K ~ dunif(%s, %s)
r0 ~ dunif(%s, %s)
theta ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
K_prior_p[1], K_prior_p[2],
r0_prior_p[1], r0_prior_p[2],
theta_prior_p[1], theta_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(y[t]) + r0 * (1 - y[t]/K)* (y[t] - theta) / K
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "allen_process.bugs")
Write the priors into a list for later reference
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r0_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
theta_prior <- function(x) dunif(x, theta_prior_p[1], theta_prior_p[2])
par_priors <- list(K = K_prior, deviance = function(x) 0 * x,
r0 = r0_prior, theta = theta_prior,
stdQ = stdQ_prior)
We define which parameters to keep track of, and set the initial values of
parameters in the transformed space used by the MCMC. We use logarithms
to maintain strictly positive values of parameters where appropriate.
jags.params=c("K","r0","theta","stdQ") # be sensible about the order here
jags.inits <- function(){
list("K"= 10 * rlnorm(1,0, 0.1),
"r0"= 1 * rlnorm(1,0, 0.1) ,
"theta"= 5 * rlnorm(1,0, 0.1) ,
"stdQ"= abs( 0.1 * rlnorm(1,0, 0.1)),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(1234)
# parallel refuses to take variables as arguments (e.g. n.iter = 1e5 works, but n.iter = n doesn't)
allen_jags <- do.call(jags.parallel, list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin,
n.burnin=n.burnin,
model.file="allen_process.bugs"))
# Run again iteratively if we haven't met the Gelman-Rubin convergence criterion
recompile(allen_jags) # required for parallel
allen_jags <- do.call(autojags,
list(object=allen_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin))
Convergence diagnostics for Allen model
R notes: this strips classes from the mcmc.list object (so that we have list of matrices; objects that reshape2::melt can handle intelligently), and then combines chains into one array. In this array each parameter is given its value at each sample from the posterior (index) for each chain.
tmp <- lapply(as.mcmc(allen_jags), as.matrix) # strip classes the hard way...
allen_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(allen_posteriors) = c("index", "variable", "value", "chain")
plot_allen_traces <- ggplot(allen_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
allen_priors <- ddply(allen_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
plot_allen_posteriors <- ggplot(allen_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=allen_priors, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=3)
Reshape the posterior parameter distribution data, transform back into original space, and calculate the mean parameters and mean function
A <- allen_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
pardist <- acast(A, index ~ variable)
bayes_coef <- apply(pardist,2, posterior.mode)
bayes_pars <- unname(c(bayes_coef["r0"], bayes_coef["K"], bayes_coef["theta"])) # parameters formatted for f
allen_f <- function(x,h,p) unname(RickerAllee(x,h, unname(p[c("r0", "K", "theta")])))
allen_means <- sapply(x_grid, f, 0, bayes_pars)
bayes_pars
[1] 0.5003 7.7115 4.6039
head(pardist)
K deviance r0 stdQ theta
170 7.898 34.41 0.6668 0.05566 3.956
171 7.745 32.76 0.7554 0.05137 4.668
172 7.577 31.66 1.0705 0.04562 4.073
173 7.859 31.26 1.5647 0.04238 5.087
174 7.821 30.67 1.7348 0.04662 5.230
175 8.003 34.90 1.4495 0.05005 5.444
Parametric Bayes based on the structurally wrong model (Ricker)
K_prior_p <- c(0.01, 40.0)
r0_prior_p <- c(0.01, 20.0)
bugs.model <-
paste(sprintf(
"model{
K ~ dunif(%s, %s)
r0 ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
K_prior_p[1], K_prior_p[2],
r0_prior_p[1], r0_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(y[t]) + r0 * (1 - y[t]/K)
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "ricker_process.bugs")
Compute prior curves
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r0_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
par_priors <- list(K = K_prior, deviance = function(x) 0 * x,
r0 = r0_prior, stdQ = stdQ_prior)
We define which parameters to keep track of, and set the initial values of
parameters in the transformed space used by the MCMC.
jags.params=c("K","r0", "stdQ")
jags.inits <- function(){
list("K"= 10 * rlnorm(1,0,.5),
"r0"= rlnorm(1,0,.5),
"stdQ"=sqrt(0.05) * rlnorm(1,0,.5),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(12345)
ricker_jags <- do.call(jags.parallel,
list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin, n.burnin=n.burnin,
model.file="ricker_process.bugs"))
recompile(ricker_jags)
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
ricker_jags <- do.call(autojags,
list(object=ricker_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin,
progress.bar="none"))
Convergence diagnostics for parametric bayes Ricker model
tmp <- lapply(as.mcmc(ricker_jags), as.matrix) # strip classes the hard way...
ricker_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(ricker_posteriors) = c("index", "variable", "value", "chain")
plot_ricker_traces <- ggplot(ricker_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
ricker_priors <- ddply(ricker_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
# plot posterior distributions
plot_ricker_posteriors <- ggplot(ricker_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=ricker_priors, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=2)
Reshape posteriors data, transform back, calculate mode and corresponding function.
A <- ricker_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
ricker_pardist <- acast(A, index ~ variable)
bayes_coef <- apply(ricker_pardist,2, posterior.mode)
ricker_bayes_pars <- unname(c(bayes_coef["r0"], bayes_coef["K"]))
ricker_f <- function(x,h,p){
sapply(x, function(x){
x <- pmax(0, x-h)
pmax(0, x * exp(p["r0"] * (1 - x / p["K"] )) )
})
}
ricker_means <- sapply(x_grid, Ricker, 0, ricker_bayes_pars[c(1,2)])
head(ricker_pardist)
K deviance r0 stdQ
170 7.727 35.73 0.29440 0.05197
171 7.949 37.93 0.18845 0.05872
172 32.183 44.54 0.02474 0.05688
173 8.149 37.59 0.12204 0.05407
174 7.460 39.01 0.38643 0.05495
175 7.361 35.54 0.23805 0.05150
ricker_bayes_pars
[1] 0.1997 7.5976
Myers Parametric Bayes
r0_prior_p <- c(.0001, 10.0)
theta_prior_p <- c(.0001, 10.0)
K_prior_p <- c(.0001, 40.0)
bugs.model <-
paste(sprintf(
"model{
r0 ~ dunif(%s, %s)
theta ~ dunif(%s, %s)
K ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
r0_prior_p[1], r0_prior_p[2],
theta_prior_p[1], theta_prior_p[2],
K_prior_p[1], K_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(r0) + theta * log(y[t]) - log(1 + pow(abs(y[t]), theta) / K)
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "myers_process.bugs")
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
theta_prior <- function(x) dunif(x, theta_prior_p[1], theta_prior_p[2])
par_priors <- list( deviance = function(x) 0 * x, K = K_prior,
r0 = r_prior, theta = theta_prior,
stdQ = stdQ_prior)
jags.params=c("r0", "theta", "K", "stdQ")
jags.inits <- function(){
list("r0"= 1 * rlnorm(1,0,.1),
"K"= 10 * rlnorm(1,0,.1),
"theta" = 1 * rlnorm(1,0,.1),
"stdQ"= sqrt(0.2) * rlnorm(1,0,.1),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(12345)
myers_jags <- do.call(jags.parallel,
list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin,
n.burnin=n.burnin,
model.file="myers_process.bugs"))
recompile(myers_jags)
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
myers_jags <- do.call(autojags,
list(myers_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin,
progress.bar="none"))
Convergence diagnostics for parametric bayes
tmp <- lapply(as.mcmc(myers_jags), as.matrix) # strip classes
myers_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(myers_posteriors) = c("index", "variable", "value", "chain")
plot_myers_traces <- ggplot(myers_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
par_prior_curves <- ddply(myers_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
plot_myers_posteriors <- ggplot(myers_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=par_prior_curves, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=3)
A <- myers_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
myers_pardist <- acast(A, index ~ variable)
bayes_coef <- apply(myers_pardist,2, posterior.mode) # much better estimates
myers_bayes_pars <- unname(c(bayes_coef[2], bayes_coef[3], bayes_coef[1]))
myers_means <- sapply(x_grid, Myer_harvest, 0, myers_bayes_pars)
myers_f <- function(x,h,p) Myer_harvest(x, h, p[c("r0", "theta", "K")])
head(myers_pardist)
K deviance r0 stdQ theta
170 20.51 37.16 1.3995 0.04136 0.9864
171 17.10 41.09 0.6076 0.05170 1.8684
172 24.40 36.33 1.1692 0.04307 1.0715
173 27.56 37.46 0.9533 0.05550 1.2000
174 22.21 34.94 0.5140 0.04203 1.8694
175 36.05 33.49 0.3089 0.03977 2.1519
myers_bayes_pars
[1] 35.4576 0.5129 23.9914
Phase-space diagram of the expected dynamics
models <- data.frame(x=x_grid,
GP=tgp_dat$y,
True=true_means,
MLE=est_means,
Ricker=ricker_means,
Allen = allen_means,
Myers = myers_means)
models <- melt(models, id="x")
# some labels
names(models) <- c("x", "method", "value")
# labels for the colorkey too
model_names = c("GP", "True", "MLE", "Ricker", "Allen", "Myers")
colorkey=cbPalette
names(colorkey) = model_names
plot_gp <- ggplot(tgp_dat) + geom_ribbon(aes(x,y,ymin=ymin,ymax=ymax), fill="gray80") +
geom_line(data=models, aes(x, value, col=method), lwd=1, alpha=0.8) +
geom_point(data=obs, aes(x,y), alpha=0.8) +
xlab(expression(X[t])) + ylab(expression(X[t+1])) +
scale_colour_manual(values=cbPalette)
print(plot_gp)
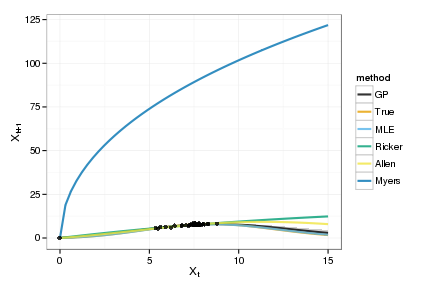
Goodness of fit
This shows only the mean predictions. For the Bayesian cases, we can instead loop over the posteriors of the parameters (or samples from the GP posterior) to get the distribution of such curves in each case.
require(MASS)
step_ahead <- function(x, f, p){
h = 0
x_predict <- sapply(x, f, h, p)
n <- length(x_predict) - 1
y <- c(x[1], x_predict[1:n])
y
}
step_ahead_posteriors <- function(x){
gp_f_at_obs <- gp_predict(gp, x, burnin=1e4, thin=300)
df_post <- melt(lapply(sample(100),
function(i){
data.frame(time = 1:length(x), stock = x,
GP = mvrnorm(1, gp_f_at_obs$Ef_posterior[,i], gp_f_at_obs$Cf_posterior[[i]]),
True = step_ahead(x,f,p),
MLE = step_ahead(x,f,est$p),
Allen = step_ahead(x, allen_f, pardist[i,]),
Ricker = step_ahead(x, ricker_f, ricker_pardist[i,]),
Myers = step_ahead(x, myers_f, myers_pardist[i,]))
}), id=c("time", "stock"))
}
df_post <- step_ahead_posteriors(x)
ggplot(df_post) + geom_point(aes(time, stock)) +
geom_line(aes(time, value, col=variable, group=interaction(L1,variable)), alpha=.1) +
scale_colour_manual(values=colorkey, guide = guide_legend(override.aes = list(alpha = 1)))
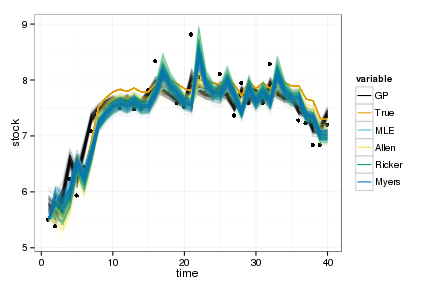
Optimal policies by value iteration
Compute the optimal policy under each model using stochastic dynamic programming. We begin with the policy based on the GP model,
# uses expected values from GP, instead of integrating over posterior
#matrices_gp <- gp_transition_matrix(gp_dat$E_Ef, gp_dat$E_Vf, x_grid, h_grid)
matrices_gp <- gp_transition_matrix(gp_dat$Ef_posterior, gp_dat$Vf_posterior, x_grid, h_grid)
opt_gp <- value_iteration(matrices_gp, x_grid, h_grid, MaxT, xT, profit, delta, reward)
Determine the optimal policy based on the allen and MLE models
matrices_true <- f_transition_matrix(f, p, x_grid, h_grid, sigma_g)
opt_true <- value_iteration(matrices_true, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
matrices_estimated <- f_transition_matrix(est$f, est$p, x_grid, h_grid, est$sigma_g)
opt_estimated <- value_iteration(matrices_estimated, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Determine the optimal policy based on Bayesian Allen model
matrices_allen <- parameter_uncertainty_SDP(allen_f, x_grid, h_grid, pardist, 4)
opt_allen <- value_iteration(matrices_allen, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Bayesian Ricker
matrices_ricker <- parameter_uncertainty_SDP(ricker_f, x_grid, h_grid, as.matrix(ricker_pardist), 3)
opt_ricker <- value_iteration(matrices_ricker, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Bayesian Myers model
matrices_myers <- parameter_uncertainty_SDP(myers_f, x_grid, h_grid, as.matrix(myers_pardist), 4)
myers_alt <- value_iteration(matrices_myers, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Assemble the data
OPT = data.frame(GP = opt_gp$D, True = opt_true$D, MLE = opt_estimated$D, Ricker = opt_ricker$D, Allen = opt_allen$D, Myers = myers_alt$D)
colorkey=cbPalette
names(colorkey) = names(OPT)
Graph of the optimal policies
policies <- melt(data.frame(stock=x_grid, sapply(OPT, function(x) x_grid[x])), id="stock")
names(policies) <- c("stock", "method", "value")
ggplot(policies, aes(stock, stock - value, color=method)) +
geom_line(lwd=1.2, alpha=0.8) + xlab("stock size") + ylab("escapement") +
scale_colour_manual(values=colorkey)
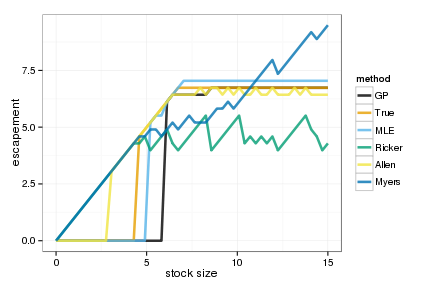
Simulate 100 realizations managed under each of the policies
sims <- lapply(OPT, function(D){
set.seed(1)
lapply(1:100, function(i)
ForwardSimulate(f, p, x_grid, h_grid, x0, D, z_g, profit=profit, OptTime=OptTime)
)
})
dat <- melt(sims, id=names(sims[[1]][[1]]))
sims_data <- data.table(dat)
setnames(sims_data, c("L1", "L2"), c("method", "reps"))
# Legend in original ordering please, not alphabetical:
sims_data$method = factor(sims_data$method, ordered=TRUE, levels=names(OPT))
ggplot(sims_data) +
geom_line(aes(time, fishstock, group=interaction(reps,method), color=method), alpha=.1) +
scale_colour_manual(values=colorkey, guide = guide_legend(override.aes = list(alpha = 1)))
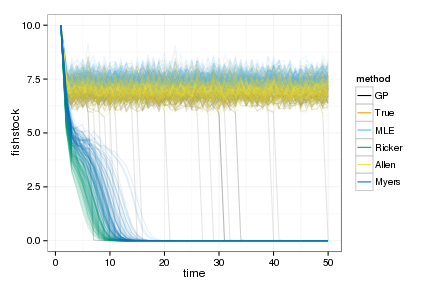
Profit <- sims_data[, sum(profit), by=c("reps", "method")]
tmp <- dcast(Profit, reps ~ method)
tmp <- tmp / tmp[,"True"]
tmp <- melt(tmp[2:dim(tmp)[2]])
actual_over_optimal <-subset(tmp, variable != "True")
allen_deviance <- posterior.mode(pardist[,'deviance'])
ricker_deviance <- posterior.mode(ricker_pardist[,'deviance'])
myers_deviance <- posterior.mode(myers_pardist[,'deviance'])
true_deviance <- 2*estf(c(p, sigma_g))
mle_deviance <- 2*estf(c(est$p, est$sigma_g))
c(allen = allen_deviance, ricker=ricker_deviance, myers=myers_deviance, true=true_deviance, mle=mle_deviance)
allen ricker myers true mle
32.93 35.92 35.46 -61.08 -889.99
cboettig/nonparametric-bayes documentation built on May 13, 2019, 2:09 p.m.
Comparison of Nonparametric Bayesian Gaussian Process estimates to standard the Parametric Bayesian approach
Plotting and knitr options, (can generally be ignored)
require(modeest)
posterior.mode <- function(x) {
mlv(x, method="shorth")$M
}
Model and parameters
Uses the model derived in citet("10.1080/10236190412331335373"), of a Ricker-like growth curve with an allee effect, defined in the pdgControl package,
f <- RickerAllee
p <- c(2, 8, 5)
K <- 10 # approx, a li'l' less
allee <- 5 # approx, a li'l' less
Various parameters defining noise dynamics, grid, and policy costs.
sigma_g <- 0.05
sigma_m <- 0.0
z_g <- function() rlnorm(1, 0, sigma_g)
z_m <- function() 1
x_grid <- seq(0, 1.5 * K, length=50)
h_grid <- x_grid
profit <- function(x,h) pmin(x, h)
delta <- 0.01
OptTime <- 50 # stationarity with unstable models is tricky thing
reward <- 0
xT <- 0
Xo <- allee+.5# observations start from
x0 <- K # simulation under policy starts from
Tobs <- 40
MaxT = 1000 # timeout for value iteration convergence
Sample Data
set.seed(1234)
#harvest <- sort(rep(seq(0, .5, length=7), 5))
x <- numeric(Tobs)
x[1] <- Xo
nz <- 1
for(t in 1:(Tobs-1))
x[t+1] = z_g() * f(x[t], h=0, p=p)
obs <- data.frame(x = c(rep(0,nz),
pmax(rep(0,Tobs-1), x[1:(Tobs-1)])),
y = c(rep(0,nz),
x[2:Tobs]))
raw_plot <- ggplot(data.frame(time = 1:Tobs, x=x), aes(time,x)) + geom_line()
raw_plot

Maximum Likelihood
set.seed(12345)
estf <- function(p){
mu <- f(obs$x,0,p)
-sum(dlnorm(obs$y, log(mu), p[4]), log=TRUE)
}
par <- c(p[1]*rlnorm(1,0,.1),
p[2]*rlnorm(1,0,.1),
p[3]*rlnorm(1,0, .1),
sigma_g * rlnorm(1,0,.1))
o <- optim(par, estf, method="L", lower=c(1e-5,1e-5,1e-5,1e-5))
f_alt <- f
p_alt <- c(as.numeric(o$par[1]), as.numeric(o$par[2]), as.numeric(o$par[3]))
sigma_g_alt <- as.numeric(o$par[4])
est <- list(f = f_alt, p = p_alt, sigma_g = sigma_g_alt, mloglik=o$value)
Mean predictions
true_means <- sapply(x_grid, f, 0, p)
est_means <- sapply(x_grid, est$f, 0, est$p)
Non-parametric Bayes
#inv gamma has mean b / (a - 1) (assuming a>1) and variance b ^ 2 / ((a - 2) * (a - 1) ^ 2) (assuming a>2)
s2.p <- c(5,5)
d.p = c(10, 1/0.1)
Estimate the Gaussian Process (nonparametric Bayesian fit)
gp <- gp_mcmc(obs$x, y=obs$y, n=1e5, s2.p = s2.p, d.p = d.p)
gp_dat <- gp_predict(gp, x_grid, burnin=1e4, thin=300)
Show traces and posteriors against priors
gp_assessment_plots <- summary_gp_mcmc(gp, burnin=1e4, thin=300)
# Summarize the GP model
tgp_dat <-
data.frame( x = x_grid,
y = gp_dat$E_Ef,
ymin = gp_dat$E_Ef - 2 * sqrt(gp_dat$E_Vf),
ymax = gp_dat$E_Ef + 2 * sqrt(gp_dat$E_Vf) )
Parametric Bayesian Models
We use the JAGS Gibbs sampler, a recent open source BUGS
implementation with an R interface that works on most platforms.
We initialize the usual MCMC parameters; see ?jags for details.
All parametric Bayesian estimates use the following basic parameters for the JAGS MCMC:
y <- x
N <- length(x);
jags.data <- list("N","y")
n.chains <- 6
n.iter <- 1e6
n.burnin <- floor(10000)
n.thin <- max(1, floor(n.chains * (n.iter - n.burnin)/1000))
n.update <- 10
We will use the same priors for process and observation noise in each model,
stdQ_prior_p <- c(1e-6, 100)
stdR_prior_p <- c(1e-6, .1)
stdQ_prior <- function(x) dunif(x, stdQ_prior_p[1], stdQ_prior_p[2])
stdR_prior <- function(x) dunif(x, stdR_prior_p[1], stdR_prior_p[2])
Parametric Bayes of correct (Allen) model
We initiate the MCMC chain (init_p) using the true values of the
parameters p from the simulation. While impossible in real data, this
gives the parametric Bayesian approach the best chance at succeeding.
y is the timeseries (recall obs has the $x_t$, $x_{t+1}$ pairs)
The actual model is defined in a model.file that contains an R function
that is automatically translated into BUGS code by R2WinBUGS. The file
defines the priors and the model. We write the file from R as follows:
K_prior_p <- c(0.01, 20.0)
r0_prior_p <- c(0.01, 6.0)
theta_prior_p <- c(0.01, 20.0)
bugs.model <-
paste(sprintf(
"model{
K ~ dunif(%s, %s)
r0 ~ dunif(%s, %s)
theta ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
K_prior_p[1], K_prior_p[2],
r0_prior_p[1], r0_prior_p[2],
theta_prior_p[1], theta_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(y[t]) + r0 * (1 - y[t]/K)* (y[t] - theta) / K
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "allen_process.bugs")
Write the priors into a list for later reference
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r0_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
theta_prior <- function(x) dunif(x, theta_prior_p[1], theta_prior_p[2])
par_priors <- list(K = K_prior, deviance = function(x) 0 * x,
r0 = r0_prior, theta = theta_prior,
stdQ = stdQ_prior)
We define which parameters to keep track of, and set the initial values of parameters in the transformed space used by the MCMC. We use logarithms to maintain strictly positive values of parameters where appropriate.
jags.params=c("K","r0","theta","stdQ") # be sensible about the order here
jags.inits <- function(){
list("K"= 10 * rlnorm(1,0, 0.1),
"r0"= 1 * rlnorm(1,0, 0.1) ,
"theta"= 5 * rlnorm(1,0, 0.1) ,
"stdQ"= abs( 0.1 * rlnorm(1,0, 0.1)),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(1234)
# parallel refuses to take variables as arguments (e.g. n.iter = 1e5 works, but n.iter = n doesn't)
allen_jags <- do.call(jags.parallel, list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin,
n.burnin=n.burnin,
model.file="allen_process.bugs"))
# Run again iteratively if we haven't met the Gelman-Rubin convergence criterion
recompile(allen_jags) # required for parallel
allen_jags <- do.call(autojags,
list(object=allen_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin))
Convergence diagnostics for Allen model
R notes: this strips classes from the mcmc.list object (so that we have list of matrices; objects that reshape2::melt can handle intelligently), and then combines chains into one array. In this array each parameter is given its value at each sample from the posterior (index) for each chain.
tmp <- lapply(as.mcmc(allen_jags), as.matrix) # strip classes the hard way...
allen_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(allen_posteriors) = c("index", "variable", "value", "chain")
plot_allen_traces <- ggplot(allen_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
allen_priors <- ddply(allen_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
plot_allen_posteriors <- ggplot(allen_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=allen_priors, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=3)
Reshape the posterior parameter distribution data, transform back into original space, and calculate the mean parameters and mean function
A <- allen_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
pardist <- acast(A, index ~ variable)
bayes_coef <- apply(pardist,2, posterior.mode)
bayes_pars <- unname(c(bayes_coef["r0"], bayes_coef["K"], bayes_coef["theta"])) # parameters formatted for f
allen_f <- function(x,h,p) unname(RickerAllee(x,h, unname(p[c("r0", "K", "theta")])))
allen_means <- sapply(x_grid, f, 0, bayes_pars)
bayes_pars
[1] 0.5003 7.7115 4.6039
head(pardist)
K deviance r0 stdQ theta
170 7.898 34.41 0.6668 0.05566 3.956
171 7.745 32.76 0.7554 0.05137 4.668
172 7.577 31.66 1.0705 0.04562 4.073
173 7.859 31.26 1.5647 0.04238 5.087
174 7.821 30.67 1.7348 0.04662 5.230
175 8.003 34.90 1.4495 0.05005 5.444
Parametric Bayes based on the structurally wrong model (Ricker)
K_prior_p <- c(0.01, 40.0)
r0_prior_p <- c(0.01, 20.0)
bugs.model <-
paste(sprintf(
"model{
K ~ dunif(%s, %s)
r0 ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
K_prior_p[1], K_prior_p[2],
r0_prior_p[1], r0_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(y[t]) + r0 * (1 - y[t]/K)
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "ricker_process.bugs")
Compute prior curves
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r0_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
par_priors <- list(K = K_prior, deviance = function(x) 0 * x,
r0 = r0_prior, stdQ = stdQ_prior)
We define which parameters to keep track of, and set the initial values of parameters in the transformed space used by the MCMC.
jags.params=c("K","r0", "stdQ")
jags.inits <- function(){
list("K"= 10 * rlnorm(1,0,.5),
"r0"= rlnorm(1,0,.5),
"stdQ"=sqrt(0.05) * rlnorm(1,0,.5),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(12345)
ricker_jags <- do.call(jags.parallel,
list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin, n.burnin=n.burnin,
model.file="ricker_process.bugs"))
recompile(ricker_jags)
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 249
Initializing model
ricker_jags <- do.call(autojags,
list(object=ricker_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin,
progress.bar="none"))
Convergence diagnostics for parametric bayes Ricker model
tmp <- lapply(as.mcmc(ricker_jags), as.matrix) # strip classes the hard way...
ricker_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(ricker_posteriors) = c("index", "variable", "value", "chain")
plot_ricker_traces <- ggplot(ricker_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
ricker_priors <- ddply(ricker_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
# plot posterior distributions
plot_ricker_posteriors <- ggplot(ricker_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=ricker_priors, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=2)
Reshape posteriors data, transform back, calculate mode and corresponding function.
A <- ricker_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
ricker_pardist <- acast(A, index ~ variable)
bayes_coef <- apply(ricker_pardist,2, posterior.mode)
ricker_bayes_pars <- unname(c(bayes_coef["r0"], bayes_coef["K"]))
ricker_f <- function(x,h,p){
sapply(x, function(x){
x <- pmax(0, x-h)
pmax(0, x * exp(p["r0"] * (1 - x / p["K"] )) )
})
}
ricker_means <- sapply(x_grid, Ricker, 0, ricker_bayes_pars[c(1,2)])
head(ricker_pardist)
K deviance r0 stdQ
170 7.727 35.73 0.29440 0.05197
171 7.949 37.93 0.18845 0.05872
172 32.183 44.54 0.02474 0.05688
173 8.149 37.59 0.12204 0.05407
174 7.460 39.01 0.38643 0.05495
175 7.361 35.54 0.23805 0.05150
ricker_bayes_pars
[1] 0.1997 7.5976
Myers Parametric Bayes
r0_prior_p <- c(.0001, 10.0)
theta_prior_p <- c(.0001, 10.0)
K_prior_p <- c(.0001, 40.0)
bugs.model <-
paste(sprintf(
"model{
r0 ~ dunif(%s, %s)
theta ~ dunif(%s, %s)
K ~ dunif(%s, %s)
stdQ ~ dunif(%s, %s)",
r0_prior_p[1], r0_prior_p[2],
theta_prior_p[1], theta_prior_p[2],
K_prior_p[1], K_prior_p[2],
stdQ_prior_p[1], stdQ_prior_p[2]),
"
iQ <- 1 / (stdQ * stdQ);
y[1] ~ dunif(0, 10)
for(t in 1:(N-1)){
mu[t] <- log(r0) + theta * log(y[t]) - log(1 + pow(abs(y[t]), theta) / K)
y[t+1] ~ dlnorm(mu[t], iQ)
}
}")
writeLines(bugs.model, "myers_process.bugs")
K_prior <- function(x) dunif(x, K_prior_p[1], K_prior_p[2])
r_prior <- function(x) dunif(x, r0_prior_p[1], r0_prior_p[2])
theta_prior <- function(x) dunif(x, theta_prior_p[1], theta_prior_p[2])
par_priors <- list( deviance = function(x) 0 * x, K = K_prior,
r0 = r_prior, theta = theta_prior,
stdQ = stdQ_prior)
jags.params=c("r0", "theta", "K", "stdQ")
jags.inits <- function(){
list("r0"= 1 * rlnorm(1,0,.1),
"K"= 10 * rlnorm(1,0,.1),
"theta" = 1 * rlnorm(1,0,.1),
"stdQ"= sqrt(0.2) * rlnorm(1,0,.1),
.RNG.name="base::Wichmann-Hill", .RNG.seed=123)
}
set.seed(12345)
myers_jags <- do.call(jags.parallel,
list(data=jags.data, inits=jags.inits,
jags.params, n.chains=n.chains,
n.iter=n.iter, n.thin=n.thin,
n.burnin=n.burnin,
model.file="myers_process.bugs"))
recompile(myers_jags)
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
Compiling model graph
Resolving undeclared variables
Allocating nodes
Graph Size: 406
Initializing model
myers_jags <- do.call(autojags,
list(myers_jags, n.update=n.update,
n.iter=n.iter, n.thin = n.thin,
progress.bar="none"))
Convergence diagnostics for parametric bayes
tmp <- lapply(as.mcmc(myers_jags), as.matrix) # strip classes
myers_posteriors <- melt(tmp, id = colnames(tmp[[1]]))
names(myers_posteriors) = c("index", "variable", "value", "chain")
plot_myers_traces <- ggplot(myers_posteriors) + geom_line(aes(index, value)) +
facet_wrap(~ variable, scale="free", ncol=1)
par_prior_curves <- ddply(myers_posteriors, "variable", function(dd){
grid <- seq(min(dd$value), max(dd$value), length = 100)
data.frame(value = grid, density = par_priors[[dd$variable[1]]](grid))
})
plot_myers_posteriors <- ggplot(myers_posteriors, aes(value)) +
stat_density(geom="path", position="identity", alpha=0.7) +
geom_line(data=par_prior_curves, aes(x=value, y=density), col="red") +
facet_wrap(~ variable, scale="free", ncol=3)
A <- myers_posteriors
A$index <- A$index + A$chain * max(A$index) # Combine samples across chains by renumbering index
myers_pardist <- acast(A, index ~ variable)
bayes_coef <- apply(myers_pardist,2, posterior.mode) # much better estimates
myers_bayes_pars <- unname(c(bayes_coef[2], bayes_coef[3], bayes_coef[1]))
myers_means <- sapply(x_grid, Myer_harvest, 0, myers_bayes_pars)
myers_f <- function(x,h,p) Myer_harvest(x, h, p[c("r0", "theta", "K")])
head(myers_pardist)
K deviance r0 stdQ theta
170 20.51 37.16 1.3995 0.04136 0.9864
171 17.10 41.09 0.6076 0.05170 1.8684
172 24.40 36.33 1.1692 0.04307 1.0715
173 27.56 37.46 0.9533 0.05550 1.2000
174 22.21 34.94 0.5140 0.04203 1.8694
175 36.05 33.49 0.3089 0.03977 2.1519
myers_bayes_pars
[1] 35.4576 0.5129 23.9914
Phase-space diagram of the expected dynamics
models <- data.frame(x=x_grid,
GP=tgp_dat$y,
True=true_means,
MLE=est_means,
Ricker=ricker_means,
Allen = allen_means,
Myers = myers_means)
models <- melt(models, id="x")
# some labels
names(models) <- c("x", "method", "value")
# labels for the colorkey too
model_names = c("GP", "True", "MLE", "Ricker", "Allen", "Myers")
colorkey=cbPalette
names(colorkey) = model_names
plot_gp <- ggplot(tgp_dat) + geom_ribbon(aes(x,y,ymin=ymin,ymax=ymax), fill="gray80") +
geom_line(data=models, aes(x, value, col=method), lwd=1, alpha=0.8) +
geom_point(data=obs, aes(x,y), alpha=0.8) +
xlab(expression(X[t])) + ylab(expression(X[t+1])) +
scale_colour_manual(values=cbPalette)
print(plot_gp)

Goodness of fit
This shows only the mean predictions. For the Bayesian cases, we can instead loop over the posteriors of the parameters (or samples from the GP posterior) to get the distribution of such curves in each case.
require(MASS)
step_ahead <- function(x, f, p){
h = 0
x_predict <- sapply(x, f, h, p)
n <- length(x_predict) - 1
y <- c(x[1], x_predict[1:n])
y
}
step_ahead_posteriors <- function(x){
gp_f_at_obs <- gp_predict(gp, x, burnin=1e4, thin=300)
df_post <- melt(lapply(sample(100),
function(i){
data.frame(time = 1:length(x), stock = x,
GP = mvrnorm(1, gp_f_at_obs$Ef_posterior[,i], gp_f_at_obs$Cf_posterior[[i]]),
True = step_ahead(x,f,p),
MLE = step_ahead(x,f,est$p),
Allen = step_ahead(x, allen_f, pardist[i,]),
Ricker = step_ahead(x, ricker_f, ricker_pardist[i,]),
Myers = step_ahead(x, myers_f, myers_pardist[i,]))
}), id=c("time", "stock"))
}
df_post <- step_ahead_posteriors(x)
ggplot(df_post) + geom_point(aes(time, stock)) +
geom_line(aes(time, value, col=variable, group=interaction(L1,variable)), alpha=.1) +
scale_colour_manual(values=colorkey, guide = guide_legend(override.aes = list(alpha = 1)))

Optimal policies by value iteration
Compute the optimal policy under each model using stochastic dynamic programming. We begin with the policy based on the GP model,
# uses expected values from GP, instead of integrating over posterior
#matrices_gp <- gp_transition_matrix(gp_dat$E_Ef, gp_dat$E_Vf, x_grid, h_grid)
matrices_gp <- gp_transition_matrix(gp_dat$Ef_posterior, gp_dat$Vf_posterior, x_grid, h_grid)
opt_gp <- value_iteration(matrices_gp, x_grid, h_grid, MaxT, xT, profit, delta, reward)
Determine the optimal policy based on the allen and MLE models
matrices_true <- f_transition_matrix(f, p, x_grid, h_grid, sigma_g)
opt_true <- value_iteration(matrices_true, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
matrices_estimated <- f_transition_matrix(est$f, est$p, x_grid, h_grid, est$sigma_g)
opt_estimated <- value_iteration(matrices_estimated, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Determine the optimal policy based on Bayesian Allen model
matrices_allen <- parameter_uncertainty_SDP(allen_f, x_grid, h_grid, pardist, 4)
opt_allen <- value_iteration(matrices_allen, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Bayesian Ricker
matrices_ricker <- parameter_uncertainty_SDP(ricker_f, x_grid, h_grid, as.matrix(ricker_pardist), 3)
opt_ricker <- value_iteration(matrices_ricker, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Bayesian Myers model
matrices_myers <- parameter_uncertainty_SDP(myers_f, x_grid, h_grid, as.matrix(myers_pardist), 4)
myers_alt <- value_iteration(matrices_myers, x_grid, h_grid, OptTime=MaxT, xT, profit, delta=delta)
Assemble the data
OPT = data.frame(GP = opt_gp$D, True = opt_true$D, MLE = opt_estimated$D, Ricker = opt_ricker$D, Allen = opt_allen$D, Myers = myers_alt$D)
colorkey=cbPalette
names(colorkey) = names(OPT)
Graph of the optimal policies
policies <- melt(data.frame(stock=x_grid, sapply(OPT, function(x) x_grid[x])), id="stock")
names(policies) <- c("stock", "method", "value")
ggplot(policies, aes(stock, stock - value, color=method)) +
geom_line(lwd=1.2, alpha=0.8) + xlab("stock size") + ylab("escapement") +
scale_colour_manual(values=colorkey)

Simulate 100 realizations managed under each of the policies
sims <- lapply(OPT, function(D){
set.seed(1)
lapply(1:100, function(i)
ForwardSimulate(f, p, x_grid, h_grid, x0, D, z_g, profit=profit, OptTime=OptTime)
)
})
dat <- melt(sims, id=names(sims[[1]][[1]]))
sims_data <- data.table(dat)
setnames(sims_data, c("L1", "L2"), c("method", "reps"))
# Legend in original ordering please, not alphabetical:
sims_data$method = factor(sims_data$method, ordered=TRUE, levels=names(OPT))
ggplot(sims_data) +
geom_line(aes(time, fishstock, group=interaction(reps,method), color=method), alpha=.1) +
scale_colour_manual(values=colorkey, guide = guide_legend(override.aes = list(alpha = 1)))

Profit <- sims_data[, sum(profit), by=c("reps", "method")]
tmp <- dcast(Profit, reps ~ method)
tmp <- tmp / tmp[,"True"]
tmp <- melt(tmp[2:dim(tmp)[2]])
actual_over_optimal <-subset(tmp, variable != "True")
allen_deviance <- posterior.mode(pardist[,'deviance'])
ricker_deviance <- posterior.mode(ricker_pardist[,'deviance'])
myers_deviance <- posterior.mode(myers_pardist[,'deviance'])
true_deviance <- 2*estf(c(p, sigma_g))
mle_deviance <- 2*estf(c(est$p, est$sigma_g))
c(allen = allen_deviance, ricker=ricker_deviance, myers=myers_deviance, true=true_deviance, mle=mle_deviance)
allen ricker myers true mle
32.93 35.92 35.46 -61.08 -889.99
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
