README.md
In ravetools: Signal and Image Processing Toolbox for Analyzing Intracranial Electroencephalography Data
ravetools
The goal of ravetools is to provide memory-efficient signal & image
processing toolbox for intracranial Electroencephalography.
Highlighted features include:
Notch filter (remove electrical line frequencies)Welch Periodogram (averaged power over frequencies)Wavelet (frequency-time decomposition)- 2D, 3D image convolution via
FFT
CT/MRI to MRI image alignment
Installation
The package is available on CRAN. To install the compiled version,
simply run:
install.packages("ravetools")
Installing the package from source requires installation of proper
compilers and some C libraries; see this
document
for details.
iEEG preprocess pipeline
This is a basic example which shows you how to preprocess an iEEG
signal. The goal here is to:
- Plot diagnostic graphs to inspect channels
- Apply Notch filters to remove electrical line noise
- Frequency-time decomposition and show the power densities
* Channel referencing is not included
1. Generate toy examples:
library(ravetools)
# Generate 20 second data at 2000 Hz
time <- seq(0, 20, by = 1 / 2000)
signal <- sin( 120 * pi * time) +
sin(time * 20*pi) +
exp(-time^2) *
cos(time * 10*pi) +
rnorm(length(time))
diagnose_channel(signal, srate = 2000)
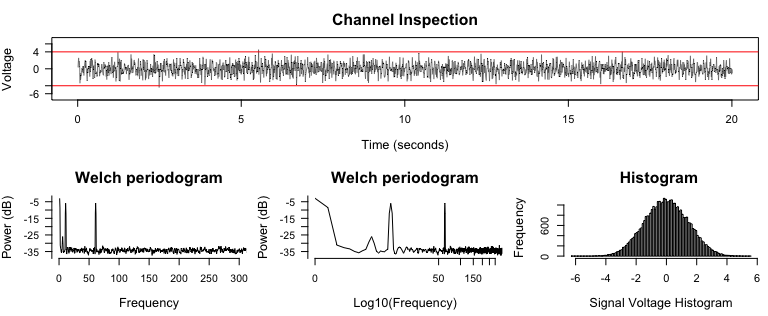
2. Apply Notch filters and inspect Periodograms
## ------- Notch filter --------
signal2 <- notch_filter(signal, sample_rate = 2000)
diagnose_channel(signal, signal2, srate = 2000,
name = c("Raw", "Filtered"))
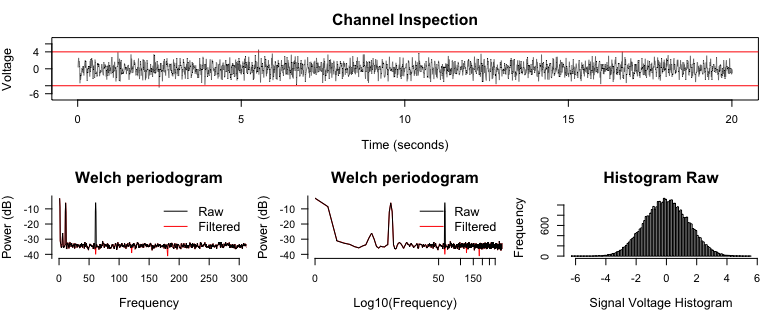
3. Frequency-time decomposition
Current version of ravetools provides two approaches: Wavelet and
Multi-taper. Wavelet uses the
Morlet wavelet and
obtains both amplitude and phase data, while Multi-taper does not
generate phase data. However, the amplitude obtained from Multi-taper
is smoother than Wavelet.
Using Wavelet:
## ---------- Wavelet -----------
coef <- morlet_wavelet(
signal2, freqs = seq(1, 100, by = 1),
srate = 2000, wave_num = c(2, 15))
amplitude <- 20 * log10(Mod(coef[]))
# For each frequency, decimate to 100 Hz
downsample_amp <- apply(amplitude, 2, decimate, q = 20)
downsample_time <- decimate(time, q = 20)
par(mfrow = c(1,1))
image(
z = downsample_amp,
x = downsample_time,
y = seq(1, 100, by = 1),
xlab = "Time (s)",
ylab = "Frequency (Hz)",
main = "Amplitude (dB)",
sub = "Wavelet at 2000 Hz, then down-sampled to 100 Hz",
col = matlab_palette()
)
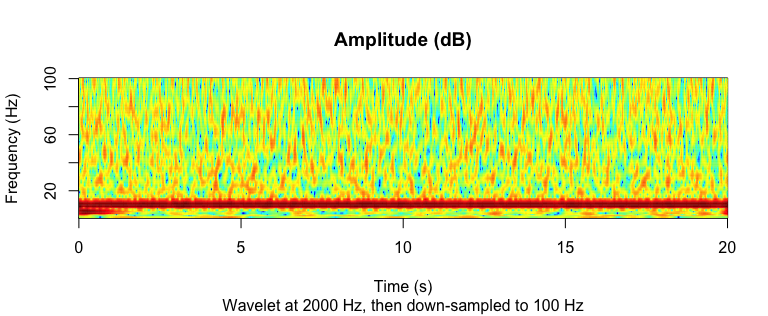
Multi-taper
Alternatively you can use Multi-tapers to obtain amplitude data. The
algorithm is modified from source code
here. Please credit
them as well if you adopt this approach.
## ---------- Multitaper -----------
res <- multitaper(
data = signal2,
fs = 2000,
frequency_range = c(1, 100),
time_bandwidth = 1.5,
window_params = c(2, 0.01),
nfft = 100
)
par(mfrow = c(1,1))
image(
x = res$time,
y = res$frequency,
z = 10 * log10(res$spec),
xlab = "Time (s)",
ylab = 'Frequency (Hz)',
col = matlab_palette(),
main = "Amplitude (dB)"
)

Image alignment
ravetools provides imaging co-registration via NiftyReg
(doi.org/10.1117/1.JMI.1.2.024003). You can align CT to MRI, or
MRI (T2) to MRI (T1). The method can be body rigid, affine, or
non-linear.
source <- system.file("extdata", "epi_t2.nii.gz", package="RNiftyReg")
target <- system.file("extdata", "flash_t1.nii.gz", package="RNiftyReg")
aligned <- register_volume(source, target, verbose = FALSE)
source_img <- aligned$source[[1]]
target_img <- aligned$target
aligned_img <- aligned$image
par(mfrow = c(2, 2), mar = c(0.1, 0.1, 3.1, 0.1))
pal <- grDevices::grey.colors(256, alpha = 1)
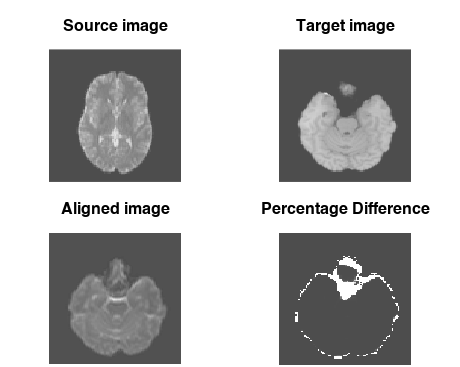
image(source_img[,,30], asp = 1, axes = FALSE,
col = pal, main = "Source image")
image(target_img[,,64], asp = 1, axes = FALSE,
col = pal, main = "Target image")
image(aligned_img[,,64], asp = 1, axes = FALSE,
col = pal, main = "Aligned image")
# bucket fill and calculate differences
aligned_img[is.nan(aligned_img) | aligned_img <= 1] <- 1
target_img[is.nan(target_img) | aligned_img <= 1] <- 1
diff <- abs(aligned_img / target_img - 1)
image(diff[,,64], asp = 1, axes = FALSE,
col = pal, main = "Percentage Difference")
References
To cite ravetools in publications use, please cite the RAVE paper from Beauchamp's lab
Magnotti, JF, and Wang, Z, and Beauchamp, MS. RAVE: comprehensive
open-source software for reproducible analysis and visualization of
intracranial EEG data. NeuroImage, 223, p.117341.
The multitaper function (MIT License) uses the script derived from
Prerau's lab. The TinyParallel script is derived from RcppParallel
package (GPL License) with TBB features removed (only use
tinythreads). The register_volume function uses NiftyReg (BSD
License) developed by CMIC at University College London, UK (its R
implementation is released under GPL license).
[1] Magnotti, JF, and Wang, Z, and Beauchamp, MS. RAVE: comprehensive
open-source software for reproducible analysis and visualization of
intracranial EEG data. NeuroImage, 223, p.117341.
[2] Prerau, Michael J, and Brown, Ritchie E, and Bianchi, Matt T, and
Ellenbogen, Jeffrey M, and Purdon, Patrick L. Sleep Neurophysiological
Dynamics Through the Lens of Multitaper Spectral Analysis. Physiology,
December 7, 2016, 60-92.
[3] Modat, M., Cash, D.M., Daga, P., Winston, G.P., Duncan, J.S. and
Ourselin, S., 2014. Global image registration using a symmetric
block-matching approach. Journal of medical imaging, 1(2), pp.024003-024003.
[4] JJ Allaire, Romain Francois, Kevin Ushey, Gregory Vandenbrouck, Marcus
Geelnard and Intel (2022). RcppParallel: Parallel Programming Tools for
'Rcpp'. R package version 5.1.5.
https://CRAN.R-project.org/package=RcppParallel
Try the ravetools package in your browser
Any scripts or data that you put into this service are public.
ravetools documentation built on Sept. 11, 2024, 9:06 p.m.
ravetools
The goal of ravetools is to provide memory-efficient signal & image
processing toolbox for intracranial Electroencephalography.
Highlighted features include:
Notch filter(remove electrical line frequencies)Welch Periodogram(averaged power over frequencies)Wavelet(frequency-time decomposition)- 2D, 3D image convolution via
FFT CT/MRItoMRIimage alignment
Installation
The package is available on CRAN. To install the compiled version,
simply run:
install.packages("ravetools")
Installing the package from source requires installation of proper
compilers and some C libraries; see this
document
for details.
iEEG preprocess pipeline
This is a basic example which shows you how to preprocess an iEEG
signal. The goal here is to:
- Plot diagnostic graphs to inspect channels
- Apply Notch filters to remove electrical line noise
- Frequency-time decomposition and show the power densities
* Channel referencing is not included
1. Generate toy examples:
library(ravetools)
# Generate 20 second data at 2000 Hz
time <- seq(0, 20, by = 1 / 2000)
signal <- sin( 120 * pi * time) +
sin(time * 20*pi) +
exp(-time^2) *
cos(time * 10*pi) +
rnorm(length(time))
diagnose_channel(signal, srate = 2000)

2. Apply Notch filters and inspect Periodograms
## ------- Notch filter --------
signal2 <- notch_filter(signal, sample_rate = 2000)
diagnose_channel(signal, signal2, srate = 2000,
name = c("Raw", "Filtered"))

3. Frequency-time decomposition
Current version of ravetools provides two approaches: Wavelet and
Multi-taper. Wavelet uses the
Morlet wavelet and
obtains both amplitude and phase data, while Multi-taper does not
generate phase data. However, the amplitude obtained from Multi-taper
is smoother than Wavelet.
Using Wavelet:
## ---------- Wavelet -----------
coef <- morlet_wavelet(
signal2, freqs = seq(1, 100, by = 1),
srate = 2000, wave_num = c(2, 15))
amplitude <- 20 * log10(Mod(coef[]))
# For each frequency, decimate to 100 Hz
downsample_amp <- apply(amplitude, 2, decimate, q = 20)
downsample_time <- decimate(time, q = 20)
par(mfrow = c(1,1))
image(
z = downsample_amp,
x = downsample_time,
y = seq(1, 100, by = 1),
xlab = "Time (s)",
ylab = "Frequency (Hz)",
main = "Amplitude (dB)",
sub = "Wavelet at 2000 Hz, then down-sampled to 100 Hz",
col = matlab_palette()
)

Multi-taper
Alternatively you can use Multi-tapers to obtain amplitude data. The
algorithm is modified from source code
here. Please credit
them as well if you adopt this approach.
## ---------- Multitaper -----------
res <- multitaper(
data = signal2,
fs = 2000,
frequency_range = c(1, 100),
time_bandwidth = 1.5,
window_params = c(2, 0.01),
nfft = 100
)
par(mfrow = c(1,1))
image(
x = res$time,
y = res$frequency,
z = 10 * log10(res$spec),
xlab = "Time (s)",
ylab = 'Frequency (Hz)',
col = matlab_palette(),
main = "Amplitude (dB)"
)

Image alignment
ravetools provides imaging co-registration via NiftyReg
(doi.org/10.1117/1.JMI.1.2.024003). You can align CT to MRI, or
MRI (T2) to MRI (T1). The method can be body rigid, affine, or
non-linear.
source <- system.file("extdata", "epi_t2.nii.gz", package="RNiftyReg")
target <- system.file("extdata", "flash_t1.nii.gz", package="RNiftyReg")
aligned <- register_volume(source, target, verbose = FALSE)
source_img <- aligned$source[[1]]
target_img <- aligned$target
aligned_img <- aligned$image
par(mfrow = c(2, 2), mar = c(0.1, 0.1, 3.1, 0.1))
pal <- grDevices::grey.colors(256, alpha = 1)
image(source_img[,,30], asp = 1, axes = FALSE,
col = pal, main = "Source image")
image(target_img[,,64], asp = 1, axes = FALSE,
col = pal, main = "Target image")
image(aligned_img[,,64], asp = 1, axes = FALSE,
col = pal, main = "Aligned image")
# bucket fill and calculate differences
aligned_img[is.nan(aligned_img) | aligned_img <= 1] <- 1
target_img[is.nan(target_img) | aligned_img <= 1] <- 1
diff <- abs(aligned_img / target_img - 1)
image(diff[,,64], asp = 1, axes = FALSE,
col = pal, main = "Percentage Difference")

References
To cite ravetools in publications use, please cite the RAVE paper from Beauchamp's lab
Magnotti, JF, and Wang, Z, and Beauchamp, MS. RAVE: comprehensive
open-source software for reproducible analysis and visualization of
intracranial EEG data. NeuroImage, 223, p.117341.
The multitaper function (MIT License) uses the script derived from
Prerau's lab. The TinyParallel script is derived from RcppParallel
package (GPL License) with TBB features removed (only use
tinythreads). The register_volume function uses NiftyReg (BSD
License) developed by CMIC at University College London, UK (its R
implementation is released under GPL license).
[1] Magnotti, JF, and Wang, Z, and Beauchamp, MS. RAVE: comprehensive
open-source software for reproducible analysis and visualization of
intracranial EEG data. NeuroImage, 223, p.117341.
[2] Prerau, Michael J, and Brown, Ritchie E, and Bianchi, Matt T, and
Ellenbogen, Jeffrey M, and Purdon, Patrick L. Sleep Neurophysiological
Dynamics Through the Lens of Multitaper Spectral Analysis. Physiology,
December 7, 2016, 60-92.
[3] Modat, M., Cash, D.M., Daga, P., Winston, G.P., Duncan, J.S. and
Ourselin, S., 2014. Global image registration using a symmetric
block-matching approach. Journal of medical imaging, 1(2), pp.024003-024003.
[4] JJ Allaire, Romain Francois, Kevin Ushey, Gregory Vandenbrouck, Marcus
Geelnard and Intel (2022). RcppParallel: Parallel Programming Tools for
'Rcpp'. R package version 5.1.5.
https://CRAN.R-project.org/package=RcppParallel
Try the ravetools package in your browser
Any scripts or data that you put into this service are public.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
