In timjmiller/wham: Woods Hole Assessment Model (WHAM)
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>"
)
wham.dir <- find.package("wham")
knitr::opts_knit$set(root.dir = file.path(wham.dir,"extdata"))
library(knitr)
library(kableExtra)
1. Background
This is the 11th WHAM example. We assume you already have wham installed and are relatively familiar with the package. If not, read the Introduction and Tutorial.
In this vignette we show how to both simulate and estimate:
- A prior distribution and random effect for catchability.
- Allow time-varying catchability as a random effect.
- Environmental covariate effect on catchability
- Different effects of the same environmental covariate on catchability and recruitment.
2. Setup
# devtools::install_github("timjmiller/wham", dependencies=TRUE)
library(wham)
library(ggplot2)
library(tidyr)
library(dplyr)
Create a directory for this analysis:
# choose a location to save output, otherwise will be saved in working directory
write.dir <- "choose/where/to/save/output"
dir.create(write.dir)
setwd(write.dir)
3. A simple operating model
Make a basic_info list of input components defining a simple default stock. We'll then pass this to prepare_wham_input and fit_wham. This is similar to example 10, but now there are two indices.
make_digifish <- function(years = 1975:2014) {
digifish = list()
digifish$ages = 1:10
digifish$years = years
na = length(digifish$ages)
ny = length(digifish$years)
digifish$n_fleets = 1
digifish$catch_cv = matrix(0.1, ny, digifish$n_fleets)
digifish$catch_Neff = matrix(200, ny, digifish$n_fleets)
digifish$n_indices = 2
digifish$index_cv = matrix(0.3, ny, digifish$n_indices)
digifish$index_Neff = matrix(100, ny, digifish$n_indices)
digifish$fracyr_indices = matrix(0.5, ny, digifish$n_indices)
digifish$index_units = rep(1, length(digifish$n_indices)) #biomass
digifish$index_paa_units = rep(2, length(digifish$n_indices)) #abundance
digifish$maturity = t(matrix(1/(1 + exp(-1*(1:na - na/2))), na, ny))
L = 100*(1-exp(-0.3*(1:na - 0)))
W = exp(-11)*L^3
nwaa = digifish$n_indices + digifish$n_fleets + 2
digifish$waa = array(NA, dim = c(nwaa, ny, na))
for(i in 1:nwaa) digifish$waa[i,,] = t(matrix(W, na, ny))
digifish$fracyr_SSB = rep(0.25,ny)
digifish$q = rep(0.3, digifish$n_indices)
digifish$F = matrix(0.2,ny, digifish$n_fleets)
digifish$selblock_pointer_fleets = t(matrix(1:digifish$n_fleets, digifish$n_fleets, ny))
digifish$selblock_pointer_indices = t(matrix(digifish$n_fleets + 1:digifish$n_indices, digifish$n_indices, ny))
return(digifish)
}
digifish = make_digifish()
Now define other components needed by prepare_wham_input (selectivity and M).
selectivity = list(model = c(rep("logistic", digifish$n_fleets),rep("logistic", digifish$n_indices)),
initial_pars = rep(list(c(5,1)), digifish$n_fleets + digifish$n_indices)) #fleet, index
M = list(initial_means = rep(0.2, length(digifish$ages)))
Here we specify that recruitment deviations are independent random effects, with no stock-recruit relationship.
NAA_re = list(N1_pars = exp(10)*exp(-(0:(length(digifish$ages)-1))*M$initial_means[1]))
NAA_re$sigma = "rec" #random about mean
NAA_re$use_steepness = 0
NAA_re$recruit_model = 2 #random effects with a constant mean
NAA_re$recruit_pars = exp(10)
4. Setting up the q parameter for the second index to have a prior distribution.
catchability = list(prior_sd = c(NA, 0.3))
Now we can make the input list with prepare_wham_input
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability)
We can then define an operating model (OM) by simulating data (and recruitment) from this input:
om = fit_wham(input, do.fit = FALSE)
#simulate data from operating model
set.seed(0101010)
newdata = om$simulate(complete=TRUE)
Now put the simulated data in an input file with all the same configuration as the operating model.
temp = input
temp$data = newdata
Fit an estimation model that is the same as the operating model (self-test).
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE, retro.silent = TRUE)
fit$mohns_rho = mohns_rho(fit)
plot_wham_output(fit)
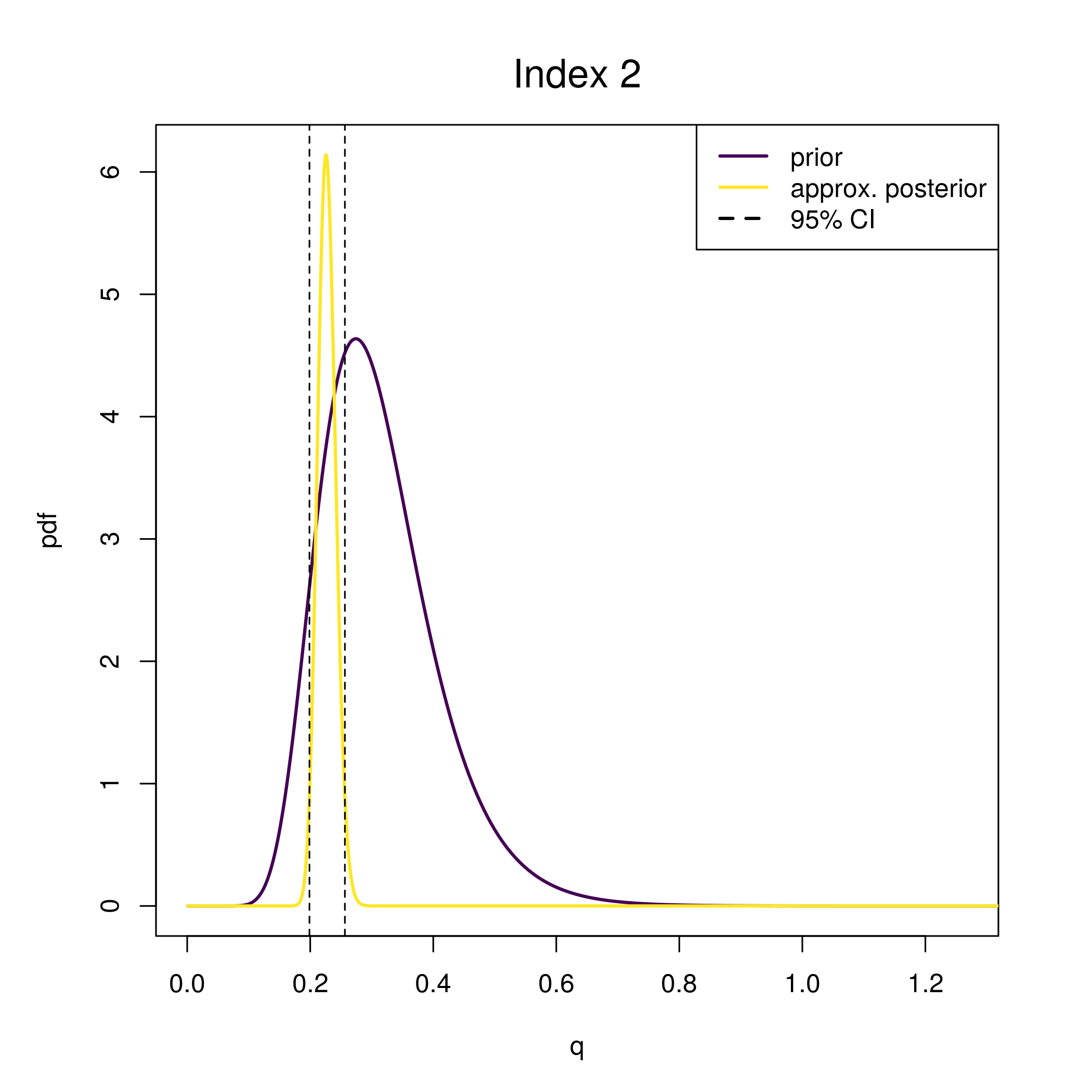
This plot of the q prior and approximate posterior is provided by plot_wham_output
wham:::plot_q_prior_post(fit)
 { width=80% }
{ width=80% }
5. Add random effects on q for first index
This will add time varying iid random effects on catchability for the first index while still keeping the prior on the second index.
catchability = list(prior_sd = c(NA, 0.3), re = c("iid", "none"))
Generate input as above.
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability)
The default initial value for the standard deviation of the random effects is 1 but we will simulate with less variation than that
input$par$q_repars[1] = log(0.2)
Now create the operating model, simulate data and fit as above
```r
om = fit_wham(input, do.fit = FALSE)
#simulate data from operating model
set.seed(0101010)
newdata = om$simulate(complete=TRUE)
#put the simulated data in an input file with all the same configuration as the operating model
temp = input
temp$data = newdata
#fit estimating model that is the same as the operating model
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)
fit$mohns_rho = mohns_rho(fit)
plot_wham_output(fit)
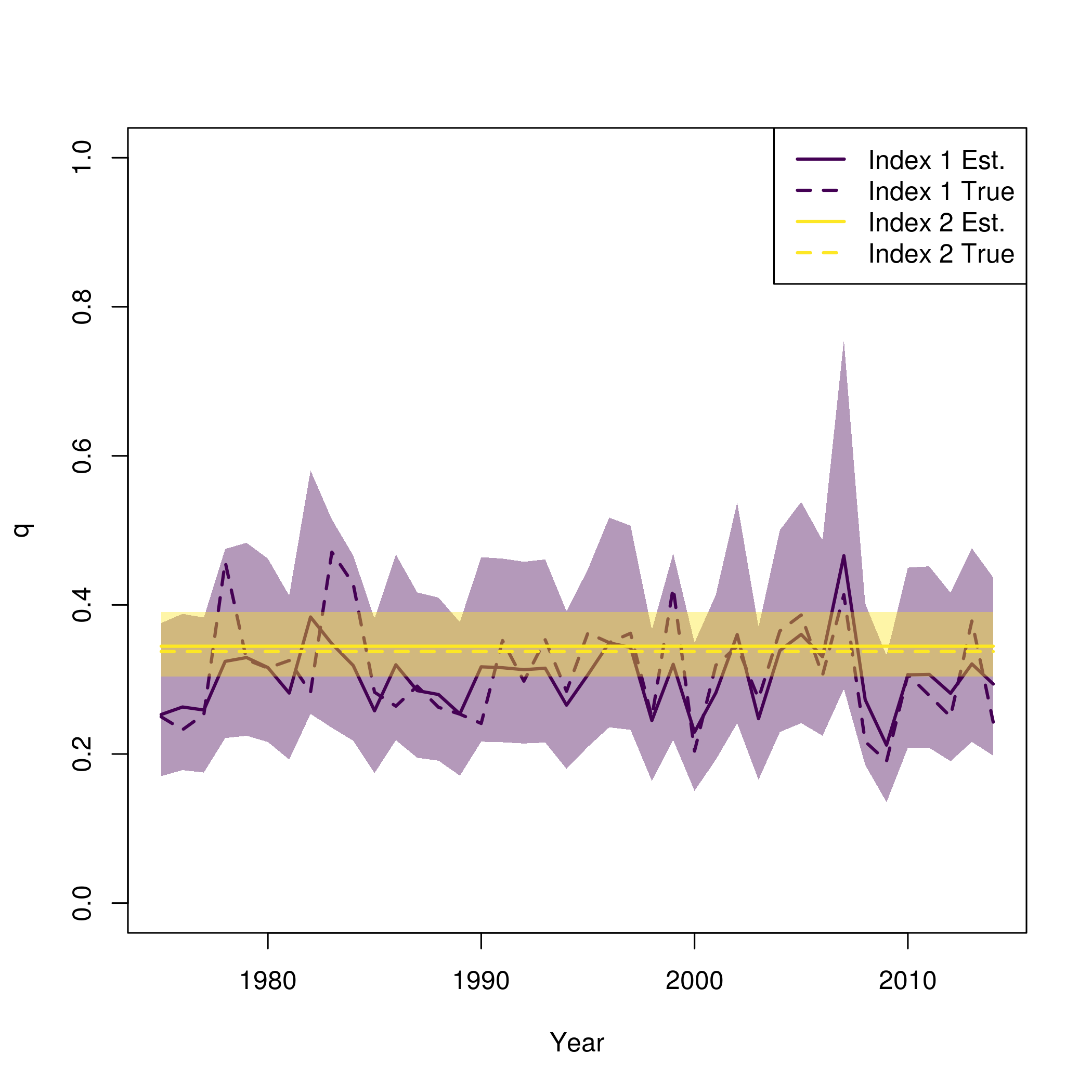
This is similar to what is provided in plot_wham_output, but with the true values from the simulation also plotted
pal = viridisLite::viridis(n=2)
plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year")
se = summary(fit$sdrep)
se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years))
for( i in 1:input$data$n_indices){
lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i])
polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i])))
polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent")
lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2)
}
legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
6. Add Environmental covariates to the model
First simulate and estimate environmental covariate processes, but no effects of population.
ecov = list(
label = c("Climate variable 1", "Climate variable 2"),
process_model = c("ar1","ar1"),
mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))),
logsigma = log(c(0.01, 0.2)),
lag = c(0,0),
years = digifish$years,
use_obs = matrix(1,length(digifish$years),2),
where = c("none","none"),
indices = list(2, NULL),
how = c(1,0))
Note that how[1] conflicts with where[1], but prepare_wham_input will use where[1].
We will not include a prior distribution for the second index, but we will keep the iid random effects on q for the first index.
catchability = list(re = c("iid", "none"))
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov)
A warning is thrown about the how[1] and where[1] conflict.
For the simulations, we need to set some values for the standard deviation and correlation parameters of the environmental covariate processes.
input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd
#cor pars are c(0.4,-0.3)
input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3)))
The same value for the standard deviation of q random effects on the first index.
input$par$q_repars[1] = log(0.2)
Now create the operating model and simulate and fit data.
om = fit_wham(input, do.fit = FALSE)
set.seed(0101010)
newdata = om$simulate(complete=TRUE)
#put the simulated data in an input file with all the same configuration as the operating model
temp = input
temp$data = newdata
#fit estimating model that is the same as the operating model
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE)
fit$mohns_rho = mohns_rho(fit)
plot_wham_output(fit)
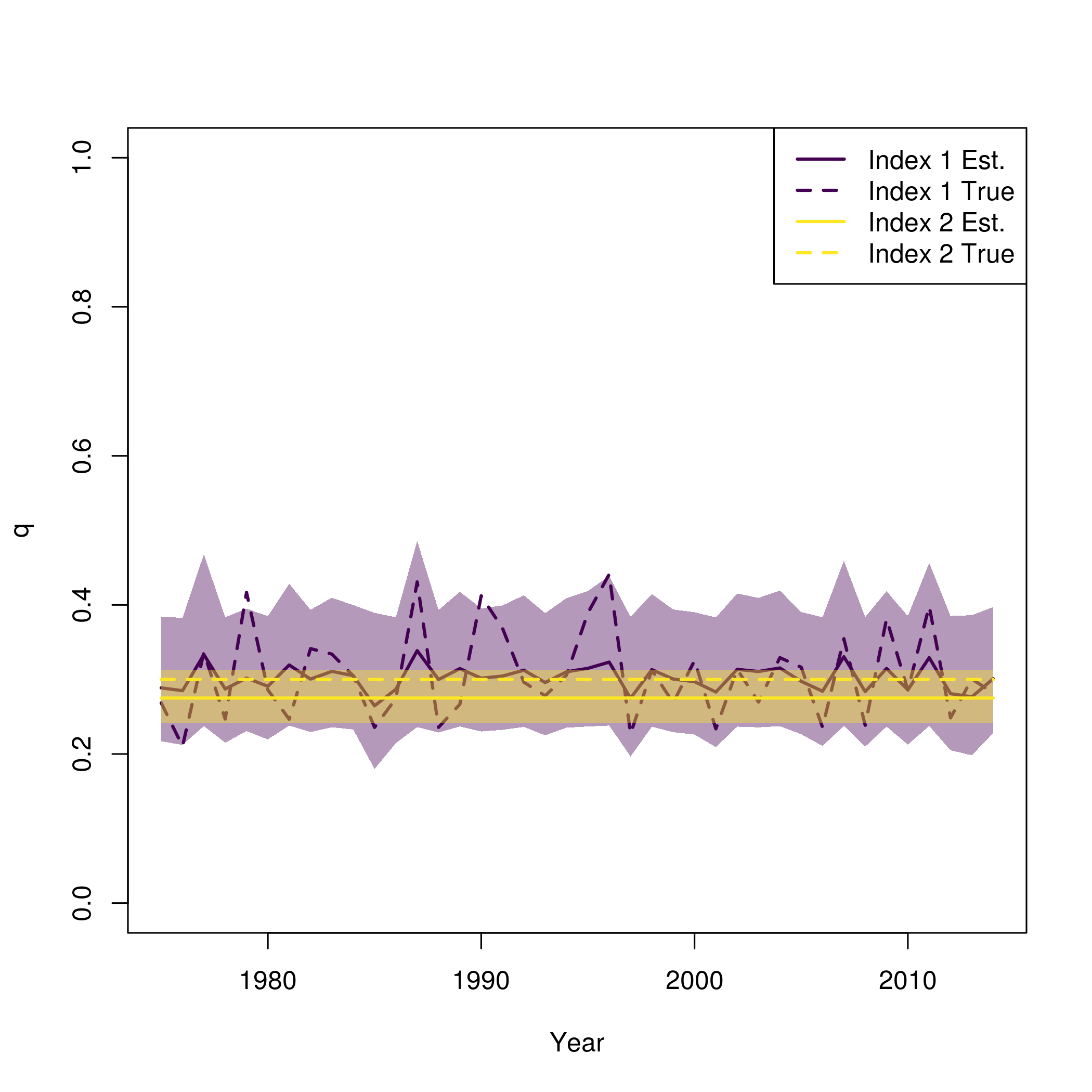
Again, this is similar to what is provided in plot_wham_output, but with the true values from the simulation also plotted
pal = viridisLite::viridis(n=2)
se = summary(fit$sdrep)
se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years))
plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year")
for( i in 1:input$data$n_indices){
lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i])
polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i])))
polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent")
lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2)
}
legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
Let's take a look at the true and estimated Environmental covariate process pararameters. Columns are the different covariates. Rows are the mean, sd, and correlation parameters, respectively.
input$par$Ecov_process_pars
fit$parList$Ecov_process_pars
Compare true and estimated (log) standard deviation of time-varying q. First row, first column.
input$par$q_repars
fit$parList$q_repars
The estimated variability in q is lower than truth, but the estimate has large standard error:
as.list(fit$sdrep, "Std")$q_repars
7. Environmental effects on catchability
Allow effects of first Environmental covariate on q for second index.
ecov = list(
label = c("Climate variable 1", "Climate variable 2"),
process_model = c("ar1","ar1"),
mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))),
logsigma = log(c(0.01, 0.2)),
lag = c(0,0),
years = digifish$years,
use_obs = matrix(1,length(digifish$years),2),
where = c("q","none"),
indices = list(2, NULL),
how = c(1,0))
As above get rid of prior on second index, add random effects on q for first index, generate input
catchability = list(re = c("iid", "none"))
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov)
how[1] no longer conflicts with where[1].
As above set parameter values for simulation.
#set sd and rho of ecov processes
input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd
#cor pars are c(0.4,-0.3)
input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3)))
#set value to simulate variation in q
input$par$q_repars[1] = log(0.2)
Also, set value for Ecov_beta effect on q (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages)
input$par$Ecov_beta[4,1,1,] = 0.5
Verify that the map argument is setting all the ``ages'' to use the same value, but only the first value is used for q or recruitment.
x = array(input$map$Ecov_beta, dim = dim(input$par$Ecov_beta))
x[4,1,1,]
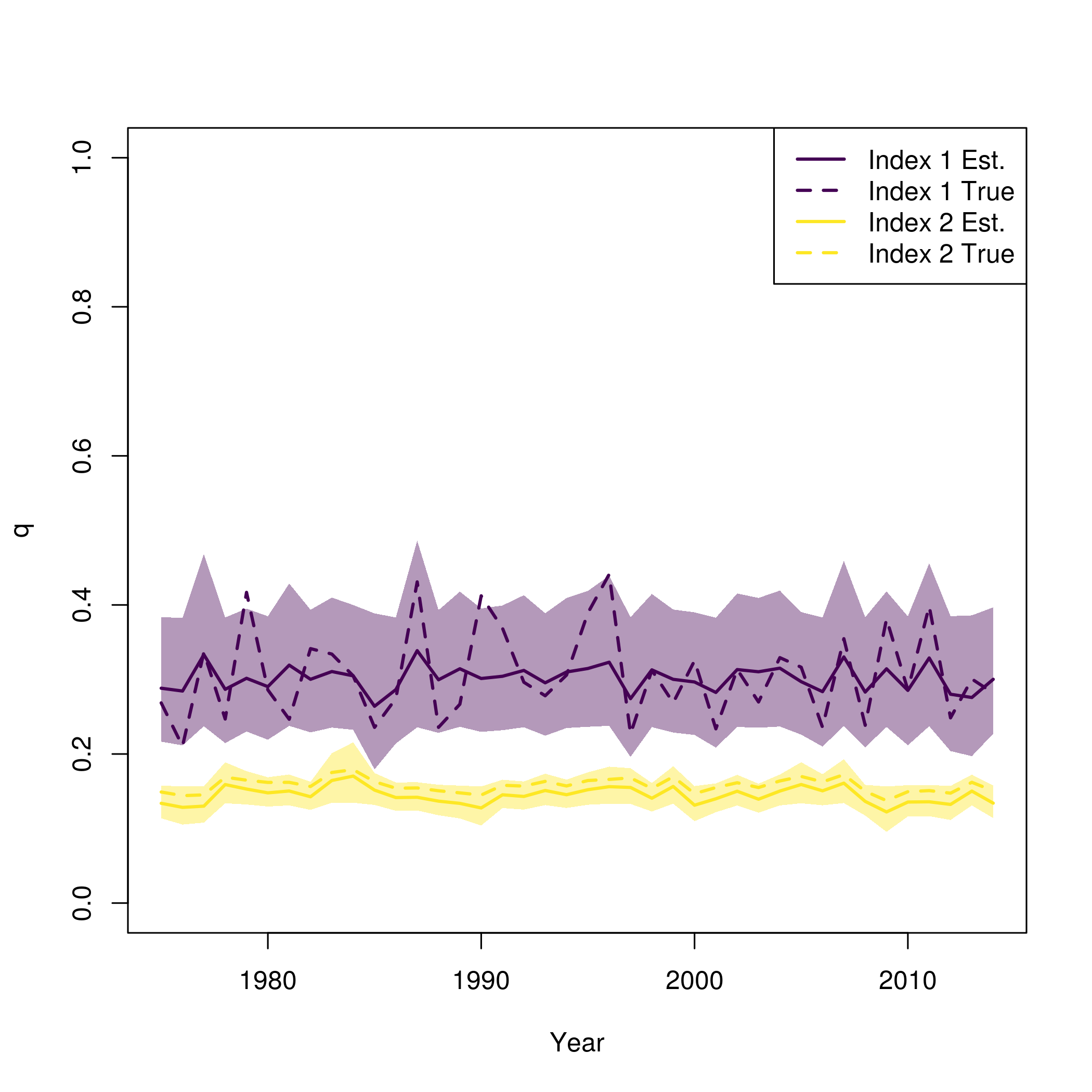
As above, create the operating model, simulate and fit data and compare true and estimated parameters.
om = fit_wham(input, do.fit = FALSE)
set.seed(0101010)
newdata = om$simulate(complete=TRUE)
#put the simulated data in an input file with all the same configuration as the operating model
temp = input
temp$data = newdata
#fit estimating model that is the same as the operating model
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE)
fit$mohns_rho = mohns_rho(fit)
plot_wham_output(fit)
#compare assumed and estimated ecov process pars
input$par$Ecov_beta[4,1,1,1]
fit$parList$Ecov_beta[4,1,1,1]
#compare assumed and estimated ecov process pars
input$par$Ecov_process_pars
fit$parList$Ecov_process_pars
#compare true and estimated time-varying q
input$par$q_repars
fit$parList$q_repars
#estimated variability in q is lower than truth, but estimate has large SE
as.list(fit$sdrep, "Std")$q_repars
pal = viridisLite::viridis(n=2)
se = summary(fit$sdrep)
se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years))
plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year")
for( i in 1:input$data$n_indices){
lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i])
polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i])))
polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent")
lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2)
}
legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
8. Different effects on q and recruitment of the same environmental covariate.
Add covariates to the model and allow effects of first covariate on q for second index AND recruitment. Then proceed as above.
ecov = list(
label = c("Climate variable 1", "Climate variable 2"),
process_model = c("ar1","ar1"),
mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))),
logsigma = log(c(0.01, 0.2)),
lag = c(0,0),
years = digifish$years,
use_obs = matrix(1,length(digifish$years),2),
where = list(c("recruit","q"),"none"),
indices = list(2, NULL),
how = c(1,0))
#get rid of prior on second index. add AR1 random effects on q for first index
catchability = list(re = c("iid", "none"))
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov)
#how[1] no longer conflicts with where[1].
#set sd and rho of ecov processes
input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd
#cor pars are c(0.4,-0.3)
input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3)))
#set value to simulate variation in q
input$par$q_repars[1] = log(0.2)
x = array(input$map$Ecov_beta, dim = dim(input$par$Ecov_beta))
x[1,1,1,] #all the ages mapped to use the same value. Only the first value is used for q or recruitment.
x[4,1,1,] #all the ages mapped to use the same value. Only the first value is used for q or recruitment.
#set value for Ecov_beta effect on q (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages)
input$par$Ecov_beta[4,1,1,] = 0.5
Also, set a value for Ecov_beta effect on recruitment (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages)
input$par$Ecov_beta[1,1,1,] = -0.5
Create the operating model, simulate and fit data, and compare true and estimated parameters.
om = fit_wham(input, do.fit = FALSE)
set.seed(0101010)
newdata = om$simulate(complete=TRUE)
#put the simulated data in an input file with all the same configuration as the operating model
temp = input
temp$data = newdata
#fit estimating model that is the same as the operating model
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE)
fit$mohns_rho = mohns_rho(fit)
plot_wham_output(fit)
#compare assumed and estimated ecov effect on q for second index
input$par$Ecov_beta[4,1,1,1]
fit$parList$Ecov_beta[4,1,1,1]
#compare assumed and estimated ecov effect on recruitment
input$par$Ecov_beta[1,1,1,1]
fit$parList$Ecov_beta[1,1,1,1]
#SE for beta parameters is large, especially for recruitment effect
as.list(fit$sdrep, "Std")$Ecov_beta[c(4,1),1,1,1]
#compare assumed and estimated ecov process pars
input$par$Ecov_process_pars
fit$parList$Ecov_process_pars
#compare true and estimated time-varying q
input$par$q_repars
fit$parList$q_repars
#estimated variability in q is lower than truth, but estimate has large SE
as.list(fit$sdrep, "Std")$q_repars
pal = viridisLite::viridis(n=2)
se = summary(fit$sdrep)
se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years))
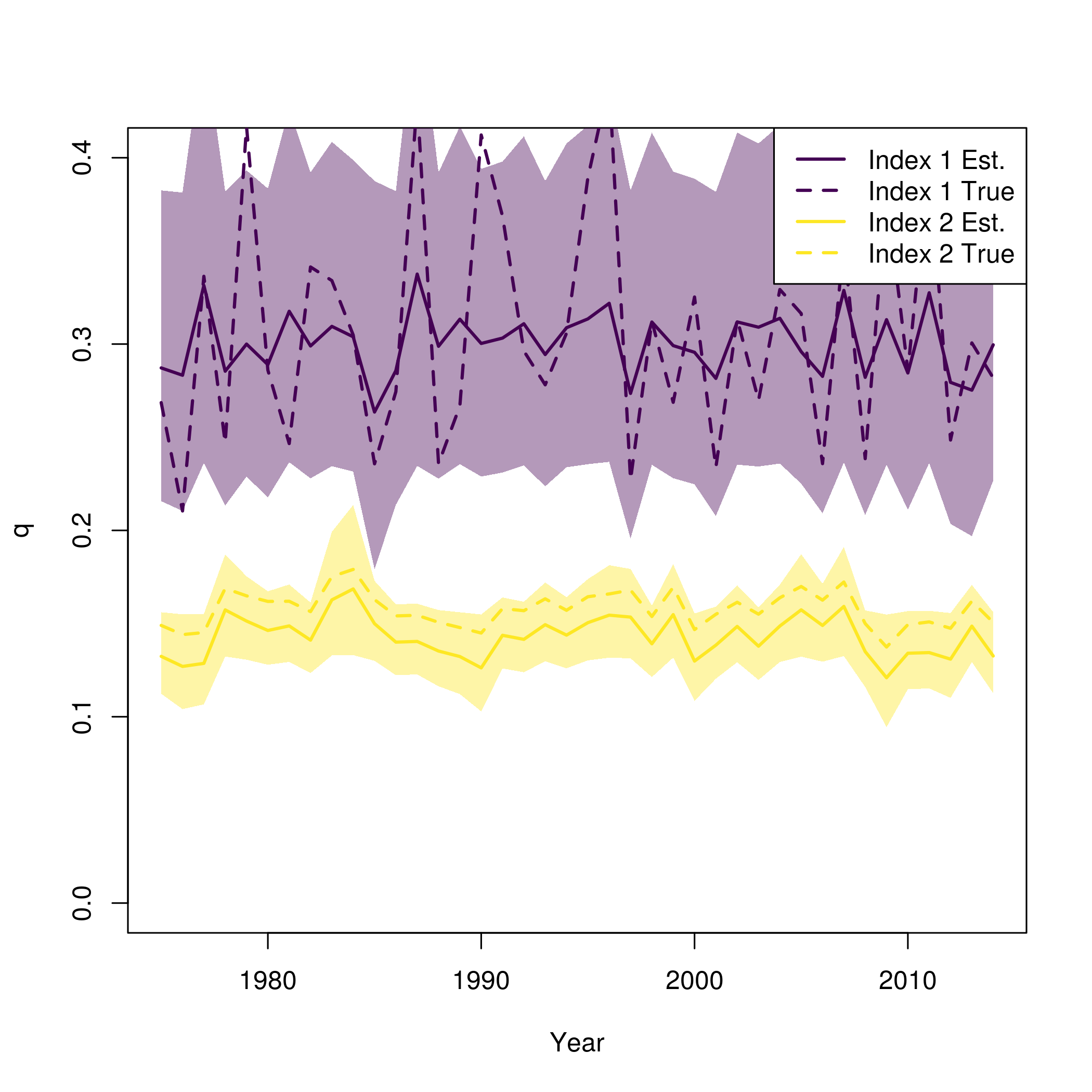
plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,0.4), ylab = "q", xlab = "Year")
for( i in 1:input$data$n_indices){
lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i])
polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i])))
polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent")
lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2)
}
legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
timjmiller/wham documentation built on April 28, 2024, 5:39 a.m.
knitr::opts_chunk$set( collapse = TRUE, comment = "#>" ) wham.dir <- find.package("wham") knitr::opts_knit$set(root.dir = file.path(wham.dir,"extdata")) library(knitr) library(kableExtra)
1. Background
This is the 11th WHAM example. We assume you already have wham installed and are relatively familiar with the package. If not, read the Introduction and Tutorial.
In this vignette we show how to both simulate and estimate:
- A prior distribution and random effect for catchability.
- Allow time-varying catchability as a random effect.
- Environmental covariate effect on catchability
- Different effects of the same environmental covariate on catchability and recruitment.
2. Setup
# devtools::install_github("timjmiller/wham", dependencies=TRUE) library(wham) library(ggplot2) library(tidyr) library(dplyr)
Create a directory for this analysis:
# choose a location to save output, otherwise will be saved in working directory write.dir <- "choose/where/to/save/output" dir.create(write.dir) setwd(write.dir)
3. A simple operating model
Make a basic_info list of input components defining a simple default stock. We'll then pass this to prepare_wham_input and fit_wham. This is similar to example 10, but now there are two indices.
make_digifish <- function(years = 1975:2014) { digifish = list() digifish$ages = 1:10 digifish$years = years na = length(digifish$ages) ny = length(digifish$years) digifish$n_fleets = 1 digifish$catch_cv = matrix(0.1, ny, digifish$n_fleets) digifish$catch_Neff = matrix(200, ny, digifish$n_fleets) digifish$n_indices = 2 digifish$index_cv = matrix(0.3, ny, digifish$n_indices) digifish$index_Neff = matrix(100, ny, digifish$n_indices) digifish$fracyr_indices = matrix(0.5, ny, digifish$n_indices) digifish$index_units = rep(1, length(digifish$n_indices)) #biomass digifish$index_paa_units = rep(2, length(digifish$n_indices)) #abundance digifish$maturity = t(matrix(1/(1 + exp(-1*(1:na - na/2))), na, ny)) L = 100*(1-exp(-0.3*(1:na - 0))) W = exp(-11)*L^3 nwaa = digifish$n_indices + digifish$n_fleets + 2 digifish$waa = array(NA, dim = c(nwaa, ny, na)) for(i in 1:nwaa) digifish$waa[i,,] = t(matrix(W, na, ny)) digifish$fracyr_SSB = rep(0.25,ny) digifish$q = rep(0.3, digifish$n_indices) digifish$F = matrix(0.2,ny, digifish$n_fleets) digifish$selblock_pointer_fleets = t(matrix(1:digifish$n_fleets, digifish$n_fleets, ny)) digifish$selblock_pointer_indices = t(matrix(digifish$n_fleets + 1:digifish$n_indices, digifish$n_indices, ny)) return(digifish) } digifish = make_digifish()
Now define other components needed by prepare_wham_input (selectivity and M).
selectivity = list(model = c(rep("logistic", digifish$n_fleets),rep("logistic", digifish$n_indices)), initial_pars = rep(list(c(5,1)), digifish$n_fleets + digifish$n_indices)) #fleet, index M = list(initial_means = rep(0.2, length(digifish$ages)))
Here we specify that recruitment deviations are independent random effects, with no stock-recruit relationship.
NAA_re = list(N1_pars = exp(10)*exp(-(0:(length(digifish$ages)-1))*M$initial_means[1])) NAA_re$sigma = "rec" #random about mean NAA_re$use_steepness = 0 NAA_re$recruit_model = 2 #random effects with a constant mean NAA_re$recruit_pars = exp(10)
4. Setting up the q parameter for the second index to have a prior distribution.
catchability = list(prior_sd = c(NA, 0.3))
Now we can make the input list with prepare_wham_input
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability)
We can then define an operating model (OM) by simulating data (and recruitment) from this input:
om = fit_wham(input, do.fit = FALSE) #simulate data from operating model set.seed(0101010) newdata = om$simulate(complete=TRUE)
Now put the simulated data in an input file with all the same configuration as the operating model.
temp = input temp$data = newdata
Fit an estimation model that is the same as the operating model (self-test).
fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE, retro.silent = TRUE) fit$mohns_rho = mohns_rho(fit) plot_wham_output(fit)
This plot of the q prior and approximate posterior is provided by plot_wham_output
wham:::plot_q_prior_post(fit)
 { width=80% }
{ width=80% }
5. Add random effects on q for first index
This will add time varying iid random effects on catchability for the first index while still keeping the prior on the second index.
catchability = list(prior_sd = c(NA, 0.3), re = c("iid", "none"))
Generate input as above.
input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability)
The default initial value for the standard deviation of the random effects is 1 but we will simulate with less variation than that
input$par$q_repars[1] = log(0.2) Now create the operating model, simulate data and fit as above ```r om = fit_wham(input, do.fit = FALSE) #simulate data from operating model set.seed(0101010) newdata = om$simulate(complete=TRUE) #put the simulated data in an input file with all the same configuration as the operating model temp = input temp$data = newdata #fit estimating model that is the same as the operating model fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE) fit$mohns_rho = mohns_rho(fit) plot_wham_output(fit)
This is similar to what is provided in plot_wham_output, but with the true values from the simulation also plotted
pal = viridisLite::viridis(n=2) plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year") se = summary(fit$sdrep) se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years)) for( i in 1:input$data$n_indices){ lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i]) polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i]))) polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent") lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2) } legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
6. Add Environmental covariates to the model
First simulate and estimate environmental covariate processes, but no effects of population.
ecov = list( label = c("Climate variable 1", "Climate variable 2"), process_model = c("ar1","ar1"), mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))), logsigma = log(c(0.01, 0.2)), lag = c(0,0), years = digifish$years, use_obs = matrix(1,length(digifish$years),2), where = c("none","none"), indices = list(2, NULL), how = c(1,0))
Note that how[1] conflicts with where[1], but prepare_wham_input will use where[1].
We will not include a prior distribution for the second index, but we will keep the iid random effects on q for the first index.
catchability = list(re = c("iid", "none")) input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov)
A warning is thrown about the how[1] and where[1] conflict.
For the simulations, we need to set some values for the standard deviation and correlation parameters of the environmental covariate processes.
input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd #cor pars are c(0.4,-0.3) input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3)))
The same value for the standard deviation of q random effects on the first index.
input$par$q_repars[1] = log(0.2)
Now create the operating model and simulate and fit data.
om = fit_wham(input, do.fit = FALSE) set.seed(0101010) newdata = om$simulate(complete=TRUE) #put the simulated data in an input file with all the same configuration as the operating model temp = input temp$data = newdata #fit estimating model that is the same as the operating model fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE) fit$mohns_rho = mohns_rho(fit) plot_wham_output(fit)
Again, this is similar to what is provided in plot_wham_output, but with the true values from the simulation also plotted
pal = viridisLite::viridis(n=2) se = summary(fit$sdrep) se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years)) plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year") for( i in 1:input$data$n_indices){ lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i]) polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i]))) polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent") lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2) } legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
Let's take a look at the true and estimated Environmental covariate process pararameters. Columns are the different covariates. Rows are the mean, sd, and correlation parameters, respectively.
input$par$Ecov_process_pars fit$parList$Ecov_process_pars
Compare true and estimated (log) standard deviation of time-varying q. First row, first column.
input$par$q_repars fit$parList$q_repars
The estimated variability in q is lower than truth, but the estimate has large standard error:
as.list(fit$sdrep, "Std")$q_repars
7. Environmental effects on catchability
Allow effects of first Environmental covariate on q for second index.
ecov = list( label = c("Climate variable 1", "Climate variable 2"), process_model = c("ar1","ar1"), mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))), logsigma = log(c(0.01, 0.2)), lag = c(0,0), years = digifish$years, use_obs = matrix(1,length(digifish$years),2), where = c("q","none"), indices = list(2, NULL), how = c(1,0))
As above get rid of prior on second index, add random effects on q for first index, generate input
catchability = list(re = c("iid", "none")) input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov)
how[1] no longer conflicts with where[1].
As above set parameter values for simulation.
#set sd and rho of ecov processes input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd #cor pars are c(0.4,-0.3) input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3))) #set value to simulate variation in q input$par$q_repars[1] = log(0.2)
Also, set value for Ecov_beta effect on q (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages)
input$par$Ecov_beta[4,1,1,] = 0.5
Verify that the map argument is setting all the ``ages'' to use the same value, but only the first value is used for q or recruitment.
x = array(input$map$Ecov_beta, dim = dim(input$par$Ecov_beta)) x[4,1,1,]
As above, create the operating model, simulate and fit data and compare true and estimated parameters.
om = fit_wham(input, do.fit = FALSE) set.seed(0101010) newdata = om$simulate(complete=TRUE) #put the simulated data in an input file with all the same configuration as the operating model temp = input temp$data = newdata #fit estimating model that is the same as the operating model fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE) fit$mohns_rho = mohns_rho(fit) plot_wham_output(fit) #compare assumed and estimated ecov process pars input$par$Ecov_beta[4,1,1,1] fit$parList$Ecov_beta[4,1,1,1] #compare assumed and estimated ecov process pars input$par$Ecov_process_pars fit$parList$Ecov_process_pars #compare true and estimated time-varying q input$par$q_repars fit$parList$q_repars #estimated variability in q is lower than truth, but estimate has large SE as.list(fit$sdrep, "Std")$q_repars pal = viridisLite::viridis(n=2) se = summary(fit$sdrep) se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years)) plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,1), ylab = "q", xlab = "Year") for( i in 1:input$data$n_indices){ lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i]) polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i]))) polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent") lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2) } legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
8. Different effects on q and recruitment of the same environmental covariate.
Add covariates to the model and allow effects of first covariate on q for second index AND recruitment. Then proceed as above.
ecov = list( label = c("Climate variable 1", "Climate variable 2"), process_model = c("ar1","ar1"), mean = cbind(rnorm(length(digifish$years)),rnorm(length(digifish$years))), logsigma = log(c(0.01, 0.2)), lag = c(0,0), years = digifish$years, use_obs = matrix(1,length(digifish$years),2), where = list(c("recruit","q"),"none"), indices = list(2, NULL), how = c(1,0)) #get rid of prior on second index. add AR1 random effects on q for first index catchability = list(re = c("iid", "none")) input = prepare_wham_input(basic_info = digifish, selectivity = selectivity, NAA_re = NAA_re, M = M, catchability = catchability, ecov = ecov) #how[1] no longer conflicts with where[1]. #set sd and rho of ecov processes input$par$Ecov_process_pars[2,] = log(c(0.1,0.2)) #sd #cor pars are c(0.4,-0.3) input$par$Ecov_process_pars[3,] = log((c(0.4,-0.3)-(-1))/(1-c(0.4,-0.3))) #set value to simulate variation in q input$par$q_repars[1] = log(0.2) x = array(input$map$Ecov_beta, dim = dim(input$par$Ecov_beta)) x[1,1,1,] #all the ages mapped to use the same value. Only the first value is used for q or recruitment. x[4,1,1,] #all the ages mapped to use the same value. Only the first value is used for q or recruitment. #set value for Ecov_beta effect on q (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages) input$par$Ecov_beta[4,1,1,] = 0.5
Also, set a value for Ecov_beta effect on recruitment (dims are n_effects (2 + n_indices, max_n_poly, n_Ecov, n_ages)
input$par$Ecov_beta[1,1,1,] = -0.5
Create the operating model, simulate and fit data, and compare true and estimated parameters.
om = fit_wham(input, do.fit = FALSE) set.seed(0101010) newdata = om$simulate(complete=TRUE) #put the simulated data in an input file with all the same configuration as the operating model temp = input temp$data = newdata #fit estimating model that is the same as the operating model fit = fit_wham(temp, do.osa = FALSE, MakeADFun.silent = TRUE)#, retro.silent = TRUE) fit$mohns_rho = mohns_rho(fit) plot_wham_output(fit) #compare assumed and estimated ecov effect on q for second index input$par$Ecov_beta[4,1,1,1] fit$parList$Ecov_beta[4,1,1,1] #compare assumed and estimated ecov effect on recruitment input$par$Ecov_beta[1,1,1,1] fit$parList$Ecov_beta[1,1,1,1] #SE for beta parameters is large, especially for recruitment effect as.list(fit$sdrep, "Std")$Ecov_beta[c(4,1),1,1,1] #compare assumed and estimated ecov process pars input$par$Ecov_process_pars fit$parList$Ecov_process_pars #compare true and estimated time-varying q input$par$q_repars fit$parList$q_repars #estimated variability in q is lower than truth, but estimate has large SE as.list(fit$sdrep, "Std")$q_repars pal = viridisLite::viridis(n=2) se = summary(fit$sdrep) se = matrix(se[rownames(se) == "logit_q_mat",2], length(fit$years)) plot(fit$years, fit$rep$q[,1], type = 'n', lwd = 2, col = pal[1], ylim = c(0,0.4), ylab = "q", xlab = "Year") for( i in 1:input$data$n_indices){ lines(fit$years, fit$rep$q[,i], lwd = 2, col = pal[i]) polyy = c(fit$rep$q[,i]*exp(-1.96*se[,i]),rev(fit$rep$q[,i]*exp(1.96*se[,i]))) polygon(c(fit$years,rev(fit$years)), polyy, col=adjustcolor(pal[i], alpha.f=0.4), border = "transparent") lines(fit$years, newdata$q[,i], lwd = 2, col = pal[i], lty = 2) } legend("topright", legend = paste0("Index ", rep(1:input$data$n_indices, each = 2), c(" Est.", " True")), lwd = 2, col = rep(pal, each = 2), lty = c(1,2))
 { width=80% }
{ width=80% }
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
