Getting Started
In threeBrain: 3D Brain Visualization
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>",
eval = FALSE
)
library(threeBrain)
threeBrain is an R package that renders FreeSurfer or AFNI/SUMA files in the modern web browsers. The goal is to provide 3D viewers that are powerful, interactive, easy-to-share.
Directory Setup
The basic setup of a typical subject folder should look like this:
Project Directory
└─Subject code
├─mri - Atlas, T1 and/or CT image files (*.mgz, transforms, ...)
├─surf - FreeSurfer surface files (pial, sulc, ...)
├─SUMA - (Optional) SUMA standard 141 surfaces (std.141.*)
└─... - Other files
If you are using FreeSurfer to generate MRI reconstruction, then you are all set! For AFNI/SUMA users, please place your SUMA folder under subject folder.
If you don't have any FreeSurfer folders, you can download a sample archive from here, choose N27-complete.zip, and extract to your download directory. Once extracted, you will have a N27 subject folder in the ~/Downloads/ directory. The directory tree looks exactly the same as described above.
In the following context, I will use ~/Downloads/N27 as an example
Generate Viewer Object
library(threeBrain)
subject_code <- "N27"
subject_path <- "~/Downloads/N27"
brain <- freesurfer_brain2(subject_path, subject_code)
print(brain)
#> Subject - N27
#> Transforms:
#>
#> - FreeSurfer TalXFM [from scanner to MNI305]:
#> [,1] [,2] [,3] [,4]
#> [1,] 0.9692 -0.0029 -0.0134 -0.1638
#> [2,] 0.0062 0.9685 0.0492 -2.0717
#> [3,] 0.0145 0.0276 0.9541 0.1361
#> [4,] 0.0000 0.0000 0.0000 1.0000
#>
#> - Torig [Voxel CRS to FreeSurfer origin, vox2ras-tkr]
#> [,1] [,2] [,3] [,4]
#> [1,] -1 0 0 128
#> [2,] 0 0 1 -128
#> [3,] 0 -1 0 128
#> [4,] 0 0 0 1
#>
#> - Norig [Voxel CRS to Scanner center, vox2ras]
#> [,1] [,2] [,3] [,4]
#> [1,] -1 0 0 128.5
#> [2,] 0 0 1 -145.5
#> [3,] 0 -1 0 146.5
#> [4,] 0 0 0 1.0
#>
#> - Scanner center relative to FreeSurfer origin
#> [1] -0.5 17.5 -18.5
#>
#> - FreeSurfer RAS to MNI305, vox2vox-MNI305
#> [,1] [,2] [,3] [,4]
#> [1,] 0.9692 -0.0029 -0.0134 0.12365
#> [2,] 0.0062 0.9685 0.0492 -18.10715
#> [3,] 0.0145 0.0276 0.9541 17.31120
#> [4,] 0.0000 0.0000 0.0000 1.00000
#> Surface information (total count 1)
#> Loading required namespace: rstudioapi
#> pial [ std.141 ]
#> Volume information (total count 1)
#> T1
If this is the first time, it might take a while to import and generate cache files.
Visualization
Visualizing the viewer is simply just one line.
brain$plot()
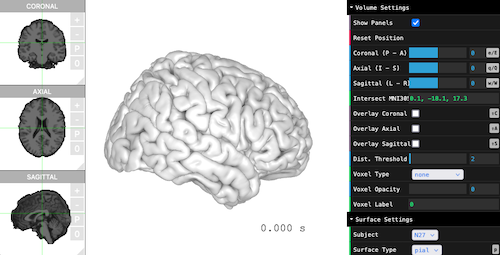
Surface, Volume, Atlas
By default, the viewers only load the following information. Only the pial surfaces are mandatory, other files are optional.
- T1 (MR images) and transforms (optional)
mri/brain.finalsurfs.mgzmri/transforms/talairach.xfmpial (mandatory) and sulc (optional)surf/*h.pial or SUMA/std.141.*h.pial[.asc|gii]surf/*h.sulc or SUMA/std.141.*h.sulc.1Daparc+aseg (optional)mri/aparc+aseg.mgz
The T1 image is automatically detected if brain.finalsurfs.mgz is missing. The following alternatives are brainmask.mgz, brainmask.auto.mgz, T1.mgz.
To load more than one surfaces, please specify the surface types when loading the viewer. The available surface types are:
pial - (default)smoothwm, white - (smoothed) white matterinflated, sphere - inflated surfacespial-outer-smoothed - ('FreeSurfer'-only) smoothed surface tightly wrapping the pial surfaceinf_200 - ('SUMA'-only) inflated pial surface
The atlas type can be selected from aparc+aseg, aparc.a2009s+aseg, aparc.DKTatlas+aseg, or aseg. To load a specific atlas, please make sure the corresponding file exists in mri/. For example, mri/aseg.mgz.
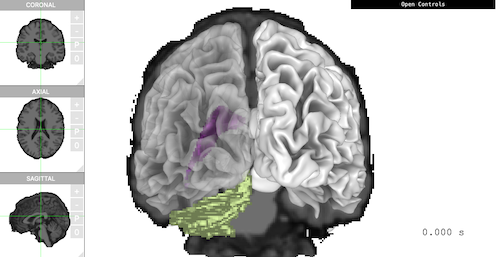
The following example loads pial and smoothwm, with aseg as atlas. The viewer shows 'Coronal' plane, smoothed white matter, 'Ventricle', and 'Cerebellum' all together in one scene.
brain <- freesurfer_brain2(
subject_path, subject_code,
surface_types = c('pial', 'smoothwm'),
atlas_types = 'aseg')
brain$plot(
controllers = list(
"Voxel Type" = "aseg",
"Voxel Label" = "4,5,6,7",
"Surface Type" = "smoothwm",
"Left Opacity" = 0.4,
"Overlay Coronal" = TRUE
),
control_display = FALSE,
camera_pos = c(0, -500, 0)
)
Try the threeBrain package in your browser
Any scripts or data that you put into this service are public.
threeBrain documentation built on July 9, 2023, 6:32 p.m.
knitr::opts_chunk$set( collapse = TRUE, comment = "#>", eval = FALSE ) library(threeBrain)
threeBrain is an R package that renders FreeSurfer or AFNI/SUMA files in the modern web browsers. The goal is to provide 3D viewers that are powerful, interactive, easy-to-share.
Directory Setup
The basic setup of a typical subject folder should look like this:
Project Directory
└─Subject code
├─mri - Atlas, T1 and/or CT image files (*.mgz, transforms, ...)
├─surf - FreeSurfer surface files (pial, sulc, ...)
├─SUMA - (Optional) SUMA standard 141 surfaces (std.141.*)
└─... - Other files
If you are using FreeSurfer to generate MRI reconstruction, then you are all set! For AFNI/SUMA users, please place your SUMA folder under subject folder.
If you don't have any FreeSurfer folders, you can download a sample archive from here, choose N27-complete.zip, and extract to your download directory. Once extracted, you will have a N27 subject folder in the ~/Downloads/ directory. The directory tree looks exactly the same as described above.
In the following context, I will use ~/Downloads/N27 as an example
Generate Viewer Object
library(threeBrain) subject_code <- "N27" subject_path <- "~/Downloads/N27" brain <- freesurfer_brain2(subject_path, subject_code) print(brain) #> Subject - N27 #> Transforms: #> #> - FreeSurfer TalXFM [from scanner to MNI305]: #> [,1] [,2] [,3] [,4] #> [1,] 0.9692 -0.0029 -0.0134 -0.1638 #> [2,] 0.0062 0.9685 0.0492 -2.0717 #> [3,] 0.0145 0.0276 0.9541 0.1361 #> [4,] 0.0000 0.0000 0.0000 1.0000 #> #> - Torig [Voxel CRS to FreeSurfer origin, vox2ras-tkr] #> [,1] [,2] [,3] [,4] #> [1,] -1 0 0 128 #> [2,] 0 0 1 -128 #> [3,] 0 -1 0 128 #> [4,] 0 0 0 1 #> #> - Norig [Voxel CRS to Scanner center, vox2ras] #> [,1] [,2] [,3] [,4] #> [1,] -1 0 0 128.5 #> [2,] 0 0 1 -145.5 #> [3,] 0 -1 0 146.5 #> [4,] 0 0 0 1.0 #> #> - Scanner center relative to FreeSurfer origin #> [1] -0.5 17.5 -18.5 #> #> - FreeSurfer RAS to MNI305, vox2vox-MNI305 #> [,1] [,2] [,3] [,4] #> [1,] 0.9692 -0.0029 -0.0134 0.12365 #> [2,] 0.0062 0.9685 0.0492 -18.10715 #> [3,] 0.0145 0.0276 0.9541 17.31120 #> [4,] 0.0000 0.0000 0.0000 1.00000 #> Surface information (total count 1) #> Loading required namespace: rstudioapi #> pial [ std.141 ] #> Volume information (total count 1) #> T1
If this is the first time, it might take a while to import and generate cache files.
Visualization
Visualizing the viewer is simply just one line.
brain$plot()

Surface, Volume, Atlas
By default, the viewers only load the following information. Only the pial surfaces are mandatory, other files are optional.
- T1 (MR images) and transforms (optional)
mri/brain.finalsurfs.mgzmri/transforms/talairach.xfmpial(mandatory) andsulc(optional)surf/*h.pialorSUMA/std.141.*h.pial[.asc|gii]surf/*h.sulcorSUMA/std.141.*h.sulc.1Daparc+aseg(optional)mri/aparc+aseg.mgz
The T1 image is automatically detected if brain.finalsurfs.mgz is missing. The following alternatives are brainmask.mgz, brainmask.auto.mgz, T1.mgz.
To load more than one surfaces, please specify the surface types when loading the viewer. The available surface types are:
pial- (default)smoothwm,white- (smoothed) white matterinflated,sphere- inflated surfacespial-outer-smoothed- ('FreeSurfer'-only) smoothed surface tightly wrapping the pial surfaceinf_200- ('SUMA'-only) inflated pial surface
The atlas type can be selected from aparc+aseg, aparc.a2009s+aseg, aparc.DKTatlas+aseg, or aseg. To load a specific atlas, please make sure the corresponding file exists in mri/. For example, mri/aseg.mgz.
The following example loads pial and smoothwm, with aseg as atlas. The viewer shows 'Coronal' plane, smoothed white matter, 'Ventricle', and 'Cerebellum' all together in one scene.
brain <- freesurfer_brain2( subject_path, subject_code, surface_types = c('pial', 'smoothwm'), atlas_types = 'aseg') brain$plot( controllers = list( "Voxel Type" = "aseg", "Voxel Label" = "4,5,6,7", "Surface Type" = "smoothwm", "Left Opacity" = 0.4, "Overlay Coronal" = TRUE ), control_display = FALSE, camera_pos = c(0, -500, 0) )

Try the threeBrain package in your browser
Any scripts or data that you put into this service are public.
Add the following code to your website.
For more information on customizing the embed code, read Embedding Snippets.
